ArciTect™ CRISPR-Cas9 Genome Editing System
ArciTect™ CRISPR-Cas9 Genome Editing System for Cell Biologists
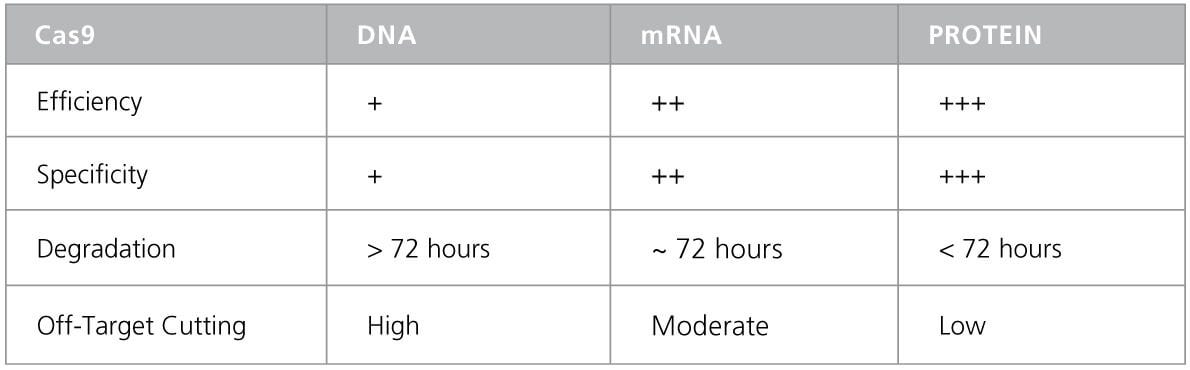
The ArciTect™ product family is a CRISPR-Cas9 genome editing system that uses ribonucleoprotein (RNP) complexes composed of purified Cas9 protein and custom synthetic guide RNA. Compared to previous technologies that utilize plasmid or mRNA-based systems, an RNP-based system enables efficient delivery and expression of CRISPR machinery in difficult-to-manipulate cells, including stem and primary cells. Once inside the cell, the RNP complexes will not induce the cellular immune response and will degrade in a timely manner to reduce off-target effects (Table 1). With validated reagents and protocols, the ArciTect™ system enables you to perform high-efficiency genome editing and generate functional gene-edited cells in your own lab.

Overcome Hurdles in Efficiency
Overcome challenges in efficient delivery and expression of CRISPR machinery using an RNP-based CRISPR-Cas9 system. ArciTect™ enables researchers to perform high-efficiency genome editing in difficult-to-manipulate cell types, including stem and primary cells.
Why Use ArciTect™?
- Maximize delivery and expression in difficult-to-manipulate cell types by using RNP complexes.
- Simplify genome editing with an integrated guide RNA design tool and cell-type-specific protocols.
- Get to your results faster with ready-to-use purified Cas9 proteins and synthetic guide RNAs.
- Minimize potential off-target cutting with timely degradation of the RNP complex.
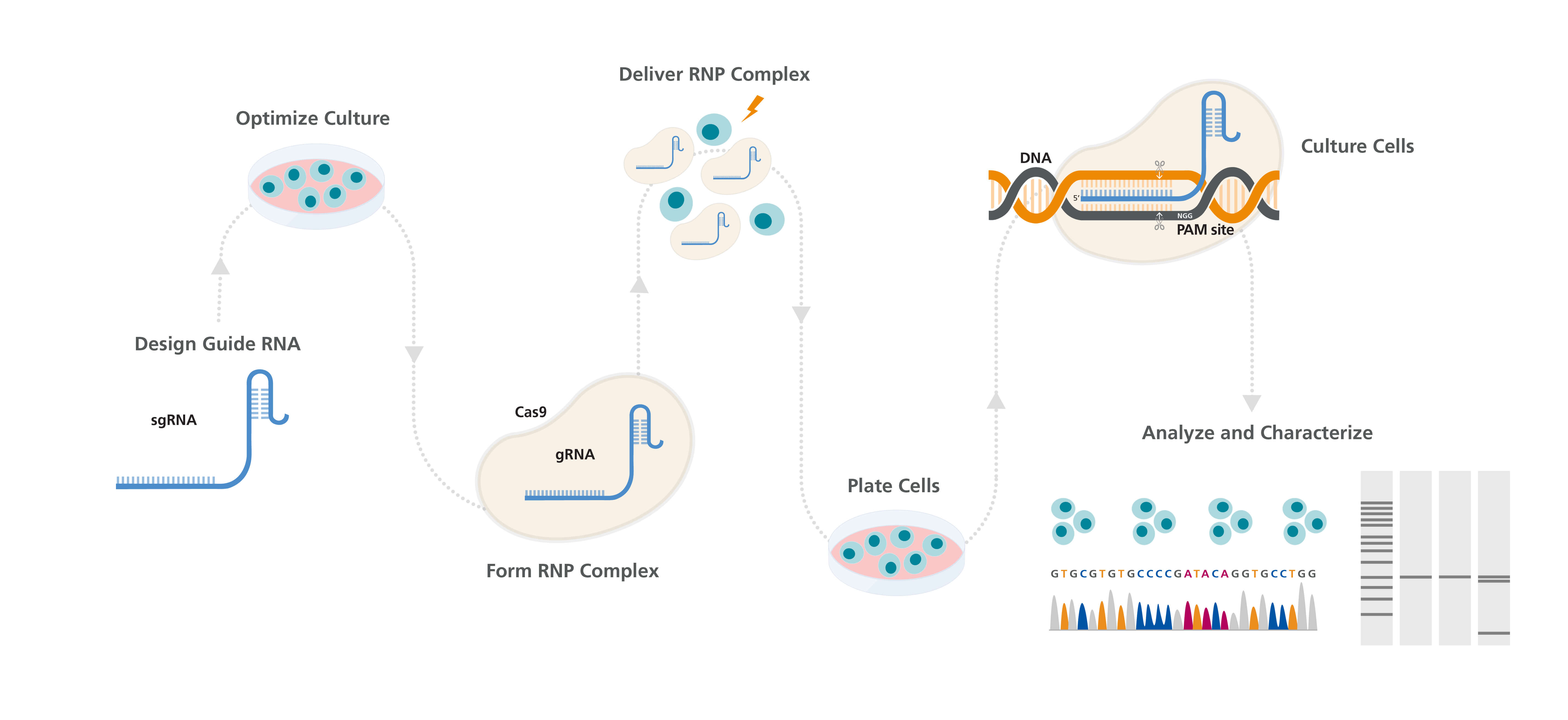
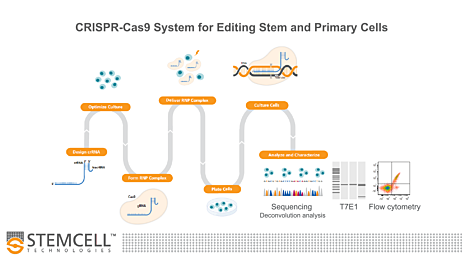
Figure 1. Genome Editing Workflow
Table 1. Comparison Between Different CRISPR Methods. 1
Genome Editing Protocols and Data
Explore step-by-step instructions for performing high-efficiency genome editing using CRISPR-Cas9 in a variety of cell lines, including stem and primary cell types. The protocols are optimized and validated with a case study and supporting data, and include important cell culture considerations and methods to evaluate editing efficiency.
Human Pluripotent Stem Cells (hPSCs)
Gene knockout and knock-in in hPSCs using electroporation and chemical transfection.
Case study: Knockout and Knock-in optimization through GFP to BFP conversion
Human Primary T Cells
Gene knockout in primary human T cells using electroporation.
Case study: Evaluation of optimal culture methods for high-efficiency TRAC knockout
CD34+ Human Hematopoietic Stem and Progenitor Cells (HSPCs)
Gene knockout in HSPCs using electroporation.
Case study: Evaluation of optimal culture methods for high-efficiency genome editing in CD34+ HSPCs
Human Intestinal Organoids
Gene knockout in human adult stem cell-derived intestinal organoids using electroporation.
Human Liver Organoids
Gene knockout in human liver organoids using electroporation.
Human Natural Killer (NK) Cells
Gene knockout in human NK cells using electroporation.
Genome Editing Products
CRISPR Design Tools
The CRISPR Guide Design Tool uses best practices and the latest computational tools to deliver the optimal CRISPR RNA (crRNA) or single guide RNA (sgRNA) sequence for every gene in the human and mouse genomes.
Scientific Resources
Wiley E-book: Genome Editing Applications
Learn about next-generation disease modeling using CRISPR, including comprehensive genome editing strategies for complex cell culture models, suggestions for optimizing experimental conditions, and troubleshooting tips.
References
- Liang X et al. (2015) Rapid and highly efficient mammalian cell engineering via Cas9 protein transfection. J Biotechnol. 208: 44-53.