Immune Response to HIV: T Cell Dysfunction, Mechanisms, and Therapies
Immunology Feature
In 1996, major advances in human immunodeficiency virus (HIV) therapy were named “Breakthrough of the Year” by Science. By combining protease inhibitors with other drugs such as azathioprine, researchers were able to substantially reduce viral loads in infected individuals. Despite this discovery being a major step forward, HIV researchers still have much to do before the disease is truly eradicated.
HIV primarily infects CD4+ T cells via their receptor, CD4, along with chemokine receptors CXCR4 and CCR51. Since CD4+ T cells are critical in orchestrating cells of adaptive immunity, such as B cells and CD8+ T cells, HIV infection impairs the immune response against the virus itself. This phenomenon explains why the immune system can’t fight HIV infection effectively. Although cells of both the innate and adaptive immune systems play important roles in controlling HIV infection, we will focus our discussion here on the interactions between HIV and T cells, and highlight a novel potential therapy against HIV infection.
T Cell Dysfunction in HIV Infection
T cells possess mechanisms to fight HIV infection (Figure 1); however, one of the hallmarks of HIV infection is T cell dysfunction, including exhaustion and progressive depletion of CD4+ T cells2,3. As shown below, both CD4+ and CD8+ T cells are rendered dysfunctional by HIV infection.
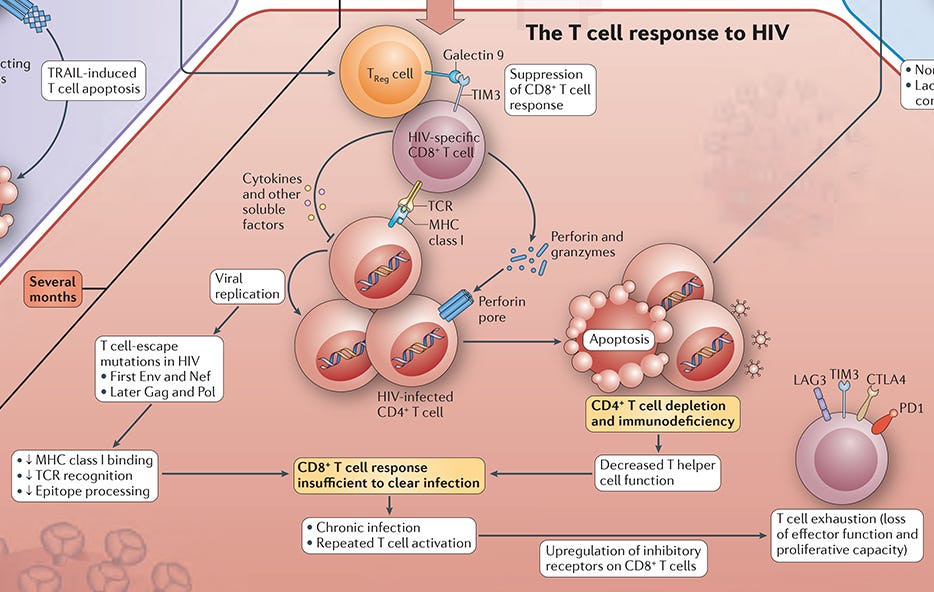
Figure 1. The T Cell Response to HIV
Adapted from the Immune Response to HIV wallchart, created by Nature Reviews Immunology in partnership with STEMCELL Technologies. Request a copy of the full Immune Response to HIV wallchart for your lab.
HIV-1 Vpu mediates HLA-C downregulation4
Cell Host & Microbe
CD8+ T cells are critical for effective antiviral responses, and viruses such as HIV-1 possess mechanisms to avoid or dampen these cells. Human leukocyte antigens (HLA), including HLA-A, HLA-B and HLA-C, present peptide antigens to CD8+ T cells, resulting in the activation of cytotoxic mechanisms to eliminate virus-infected cells. HIV-1 can cause the downregulation of HLA-A and HLA-B using the viral protein Nef. This study by Apps et al. showed that primary HIV-1 clones also downregulate HLA-C using the viral protein Vpu. As a result, HIV-1 escapes detection and elimination by CD8+ T cells to ensure its persistence.
Cell separation: EasySep™ Human CD4 Positive Selection Kit II
Upregulation of CTLA-4 by HIV-specific CD4+ T cells correlates with disease progression and defines a reversible immune dysfunction5
Nature Immunology
In chronic viral infection such as HIV infection, T cells can become exhausted due to ongoing antigen exposure3. Exhaustion is associated with reduced proliferation and cytokine production, leading to poor control of the invading virus, and is also associated with the expression of inhibitory receptors such as cytotoxic T lymphocyte-associated protein 4 (CTLA-4). Mechanistically, CTLA-4 is a co-inhibitory receptor that binds B7-1 (CD80) and B7-2 (CD86) to antagonize CD28 co-stimulation and inhibit T cell activation. In this paper, Kaufmann et al. found that CTLA-4 expression is increased in HIV-specific CD4+ T cells, and that this elevated CTLA-4 expression is associated with disease progression. Furthermore, in vitro blockade of CTLA-4 restores HIV-specific CD4+ T cell function. This study suggests that CTLA-4 blockade can potentially be used to reverse HIV-induced T cell exhaustion.
Cell separation: RosetteSep™ Human CD8 Depletion Cocktail
Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF6
Nature Medicine
CD8+ T cells also become exhausted during HIV infection, resulting in reduced proliferative capacity and ineffective antiviral responses. Programmed Death-1 (PD-1) is another key marker associated with T cell exhaustion. In 2010, the Haining lab showed that HIV-specific CD8+ T cells express PD-1 and upregulate genes associated with PD-1 signaling. Mechanistically, PD-1 upregulates basic leucine transcription factor, ATF-like (BATF) to reduce T cell proliferation and cytokine secretion. In addition to PD-1, Mueller et al. have previously shown that CD8+ T cells from HIV-infected individuals have increased susceptibility to CD95/Fas-induced apoptosis7. Overall, these studies suggest that therapies to restore CD8+ T cell function, including PD-1 blockade, may be useful to reverse T cell exhaustion in HIV infection.
Cell separation: RosetteSep™ Human CD8 Depletion Cocktail, RosetteSep™ Human CD8+ T Cell Enrichment Cocktail and RosetteSep™ Human Monocyte Enrichment Cocktail
Molecular Mechanisms to Control HIV Infection
Host cells can produce antiviral factors in an effort to eliminate invading HIV1. Although HIV can express its own factors to counteract these antiviral mechanisms, immune cell-intrinsic factors, including A3G, MX2 and RIG-I, which are discussed below, still play important roles in combating HIV infection.
The antiviral factor APOBEC3G enhances the recognition of HIV-infected primary T cells by natural killer cells8
Nature Immunology
HIV-infected T cells utilize the antiviral factor APOBEC3G (A3G), a cytidine deaminase that causes guanosine-to-adenosine hypermutation in HIV, resulting in inactivation of the virus. Interestingly, Norman et al. found that A3G also upregulates the expression of NK cell-activating ligands, NKG2D ligand, and thus enhances NK cell cytotoxicity towards HIV-infected T cells. This study describes how NK cells can detect HIV-infected T cells to help eliminate them.
Cell separation: EasySep™ Human CD56 Positive Selection Kit
MX2 is an interferon-induced inhibitor of HIV-1 infection9
Nature Letter
Type-I interferon (IFN), which includes IFN-ɑ and IFN-β, is critical for antiviral responses. Here, the Bieniasz lab identified that myxovirus resistance 2 (MX2) is an integral part of the IFN-ɑ-induced antiviral response to HIV-1. MX2 is expressed in macrophages and CD4+ T cells, among other cell types, and its expression can be induced by exposure to IFN-ɑ. Mechanistically, this study suggests that MX2 inhibits the ability of HIV-1 to import its DNA into the nuclei of host cells.
Cell separation: RosetteSep™ Human CD4+ T Cell Enrichment Cocktail
Stimulating the RIG-I pathway to kill cells in the latent HIV reservoir following viral reactivation10
Nature Medicine
This study highlights the importance of the retinoic acid-inducible gene I (RIG-I) pathway in the fight against HIV. RIG-I is a cytosolic pattern recognition receptor that recognizes intracellular dsRNA to activate antiviral signaling. HIV proviruses can remain latently hidden in host CD4+ T cells, and Li et al. suggested that activation of RIG-I pathway using acitretin, a drug currently used for the treatment of psoriasis, may help to eliminate latent HIV. Acitretin administration decreases the proviral DNA found in host CD4+ T cells and promotes the apoptosis of HIV-infected cells, thereby reducing the latent HIV reservoir in infected individuals. This article illustrates the potential of modulating immune cell-intrinsic pathways for therapeutic purposes.
Cell separation: EasySep™ Human CD4+ T Cell Isolation Kit
Novel Therapies Against HIV
The development of antiretroviral therapy has resulted in the ability to effectively treat HIV infection and significantly reduce HIV-associated deaths11; however, researchers remain hard at work to address several key challenges, including viral latency and persistence despite seemingly successful treatment.
Directed evolution of a recombinase that excises the provirus of most HIV-1 primary isolates with high specificity12
Nature Biotechnology
As mentioned above, HIV can remain latently integrated in host CD4+ T cells for a prolonged time. In this Nature Biotechnology article, Karpinski et al. developed a recombinase, Brec1, that site-specifically recognizes a 34-bp DNA sequence that can be found in a large proportion of the clinically relevant HIV-1 strains and subtypes. They showed that Brec1 can effectively and safely remove integrated HIV-1 provirus from infected cells in vitro and in vivo in humanized mice. This study suggests that Brec1 may be developed as a new potential therapy targeting latent HIV-1 infection.
Cell separation: EasySep™ Human CD4+ T Cell Isolation Kit, EasySep™ Human Cord Blood CD34 Positive Selection Kit, RoboSep™ and StemSpan™ CC110
Prevention is another key research focus that remains to be addressed. Although oral antiretroviral prophylaxis13 can be effective for at-risk populations, adherence to the prophylactic regimen is a key variable, and millions of individuals still live with HIV infection. Perhaps the notoriously elusive development of an effective HIV vaccine will be the key for the complete eradication of this virus.
Explore More Immunology Features
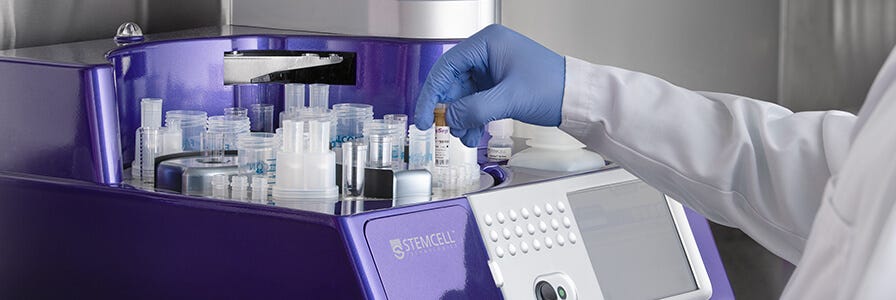
RoboSep™: Automated Cell Separation for HIV Research
HIV research often requires cell isolation procedures that minimize sample handling to avoid cross contamination of samples and exposure to the virus. RoboSep™ is a powerful instrument for rapidly isolating purified immune cells affected by HIV with minimum human manipulation.
RoboSep™ fully automates cell isolation procedures by using a robotic pipettor to perform all cell labeling and magnetic separation steps in a column-free system. The instrument can fit into a standard laminar flow hood for sterile operation and uses disposable plastic-ware tips—instead of multi-use columns and tubing—to ensure safe handling of blood products and avoid sample cross contamination.
Resources for Your HIV Research
Tools for Studying Immune Responses to Viral Infections
Learn about the immune response to viruses. Find out how obtaining the right immune cell populations and being able to analyze them effectively is critical for this research area.
Therapeutics Development for Viral Diseases
Explore technologies, applications, and research tools for developing and assessing therapeutics to combat viral diseases such as COVID-19.
References
- Barre-Sinoussi F et al. (2013) Nature Rev Microbiol 11: 877-883.
- Okoye AA et al. (2013) Immunol Rev 254(1): 54-64.
- Wherry EJ et al. (2015) Nat Rev Immunol 15(8):486-499.
- Apps R et al. (2016) Cell Host Microbe 19(5): 686-695.
- Kaufmann DE et al. (2007) Nat Immunol 8(11): 1246-1254.
- Quigley M et al. (2010) Nat Med 16(10): 1147-1151.
- Mueller YM et al. (2001) Immunity 15(6): 871-882.
- Norman JM et al. (2011) Nat Immunol 12(10): 975-983.
- Kane M et al. (2013) Nature 502(7472): 563-536.
- Li P et al. (2016) Nat Med 22(7): 807-811.
- Broder S (2011) Antiviral Res 85(1): 1.
- Karpinski J et al. (2016) Nat Biotechnol 34(4): 401-409.
- Grant RM et al (2010). N Engl J Med 363: 2587-2599.