Cell Separation and Cell Isolation Methods
There are multiple methods that can be used when performing cell isolation. Below we will discuss several cell separation techniques, including their advantages and limitations.
The cell separation method you choose typically depends on what you intend to use the isolated cells for, and the choice may involve a trade-off. For example, if you need very pure cells, you will likely choose a method with high purity but that may result in lower yield.
Consider these parameters when deciding which cell separation method to use >

Due to its speed and simplicity, immunomagnetic cell separation is one of the most commonly used methods by which scientists isolate highly purified populations of specific cell subsets. Immunomagnetic cell separation has several advantages, including:
- High purity
- Fast protocols
- Ease of use
- Low equipment cost
- Many cells can be isolated at once
- Potential for automation
- High cell viability
Both positive and negative selection can be performed using magnetic cell isolation methods. When a positive selection is performed, the supernatant can be discarded and the magnetically-labeled cells of interest remain immobilized until removed from the electromagnetic field. When a negative selection is performed, the desired cells are located in the supernatant.
Fluorescence-activated Cell Sorting
Fluorescence-activated cell sorting (FACS) is a method that uses flow cytometry and fluorescent probes to sort heterogeneous mixtures of cells. Fluorophore-tagged antibodies bind to epitopes on specific antigens on the target cells within a single-cell suspension. After tagging, the flow cytometer focuses the cell suspension into a uniform stream of single cells. This stream is then passed through a set of lasers that excites the cell-bound fluorophores, causing light scattering and fluorescent emissions. Based on the wavelengths produced by the laser excitation, the resulting photon signals are converted into a proportional number of electronic pulses that assign a charge to the droplet that is formed around the cell. As each droplet falls between the deflection plates, its charge causes the droplet to either be deflected into collection tubes or fall into the waste chamber.
Immunomagnetic cell sorting is a much faster and simpler procedure than FACS, and is often the preferred cell isolation method for common cell types. FACS has several advantages over immunomagnetic cell sorting including the ability to:
- Sort single cells
- Isolate cells based on intracellular markers (e.g. GFP)
- Isolate cells based on surface marker expression levels
- Sort complex cell types with multiple markers at higher purity
Pre-enrich samples prior to FACS
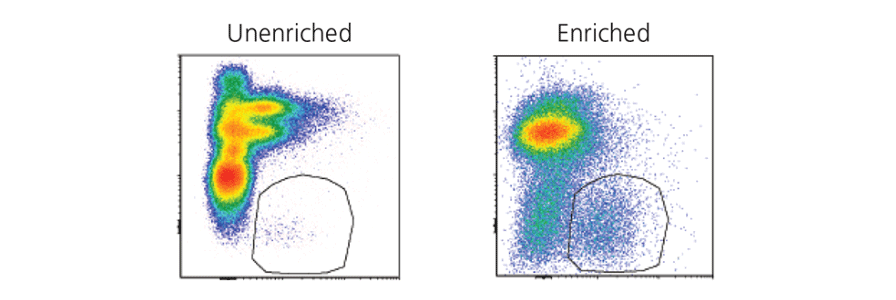
Isolating rare cell types by FACS can be time consuming, expensive and can result in low cell recovery. Researchers can pre-enrich their samples for target cells using immunomagnetic cell separation to reduce the sort time and improve purity and recovery. Read our case study:
Pre-enrichment of innate lymphoid cells (ILCs) prior to flow sorting
See how pre-enrichment using immunomagnetic cell separation can be used to save sorting time and maximize yield for ILCs, a rare cell type.
In addition to ILCs, researchers may choose to pre-enrich for other cell types, including CD4+ T cells, CD8+ T cells, B cells, or dendritic cells, prior to sorting for more specific or rare cell subsets.
View fast negative selection kits that are ideal for pre-enrichment >
Considerations for FACS gating
In flow cytometry, gates are value limits that allow you to analyze a subset of data from the larger data set. In FACS, these boundaries allow you to analyze cells with common characteristics and allows you to distinguish your cells of interest from other populations. Using an optimal gating strategy is an important factor in ensuring accurate results. For those new to flow cytometry, read the following technical tips to optimize your gating strategy before sorting your samples.
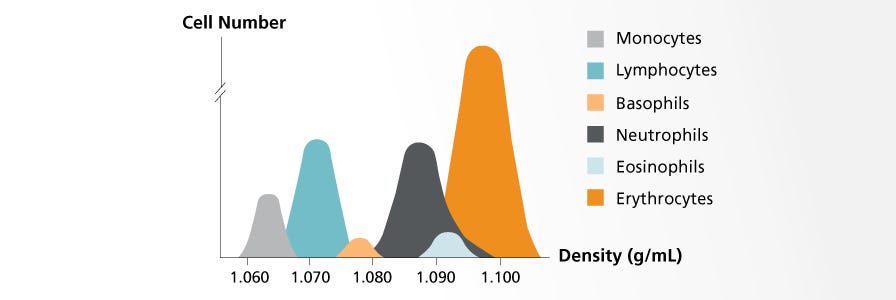
Density gradient centrifugation relies on the varying densities of cells within a heterogeneous sample. The sample is layered on top of a density gradient medium before being centrifuged. During centrifugation, each cell type will sediment to its isopycnic point, which is the place in the medium gradient where the density of the cells and medium are equal.
Common applications include the fractionation of peripheral blood mononuclear cells, exclusion of dead cells from a cell culture, and separation of plasma from blood cells.
There are several types of density gradient media, each with unique properties that render them ideal for different purposes. The following are examples of the most well-known types:
- Lymphoprep™, Lympholyte®, and Ficoll-Paque® are similar media that consist of saccharides and sodium diatrizoate; they have a density of 1.077 g/mL. These media are commonly used to isolate mononuclear cells from peripheral blood, cord blood, and bone marrow. See our comparison data >
- Percoll® (density: 1.131 g/mL) consists of colloidal silica particles coated with polyvinylpyrrolidone (PVP) and is widely used to separate cells, organelles, viruses, and other subcellular particles.
- OptiPrep™ is a medium consisting of iodixanol in water that is used to isolate viruses, organelles, macromolecules, and cells.
Density gradient centrifugation is an inexpensive cell separation technique but has limited specificity, low purity, and low throughput. In addition, even though it is a common laboratory technique, density gradient centrifugation can be a slow and laborious process that is difficult to master. Scientists typically need to carefully layer their sample over the density gradient medium, centrifuge for 30 minutes without brakes, then carefully harvest and wash the appropriate layer of cells. Technologies like SepMate™ make this method easier and faster. SepMate ™ is a specialized tube that allows users to quickly layer blood over the density gradient medium, prevents the layers from mixing and facilitates fast and easy harvesting of the target cells. With SepMate™, cells can be obtained in as little as 15 minutes.
How to use SepMate™ to simplify PBMC isolation from whole blood by density gradient centrifugation
Immunodensity Cell Separation
Immunodensity cell separation, also referred to as erythrocyte rosetting, is a negative selection method that uses a combination of antibody-based labeling and density gradient centrifugation. With this method, antibodies are added to a whole blood sample, labeling the unwanted cells and cross-linking them to red blood cells. This results in the formation of complexes called immunorosettes that are much denser than the mononuclear cells being isolated. During centrifugation, the unwanted cells pellet with the red blood cells, leaving the target cells in a layer above the density medium.
Immunodensity cell separation doesn’t require any specialized equipment beyond a centrifuge, can be easily incorporated into established density gradient centrifugation protocols, and can be used to isolate specific cell subsets directly from whole blood. However, the technique is limited to negative selection, relies on the operator’s blood sample layering technique, and requires a high concentration of red blood cells in the starting sample.
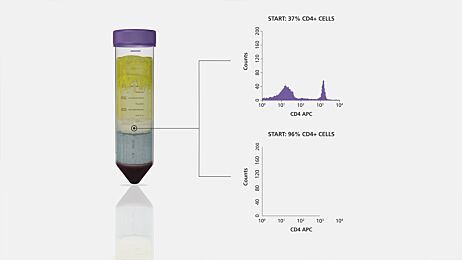
RosetteSep™ is an example of a commercially available immunodensity cell separation reagent (Figure 1). RosetteSep™ can be combined with SepMate™ PBMC isolation tubes for even faster and easier immunodensity cell separation.
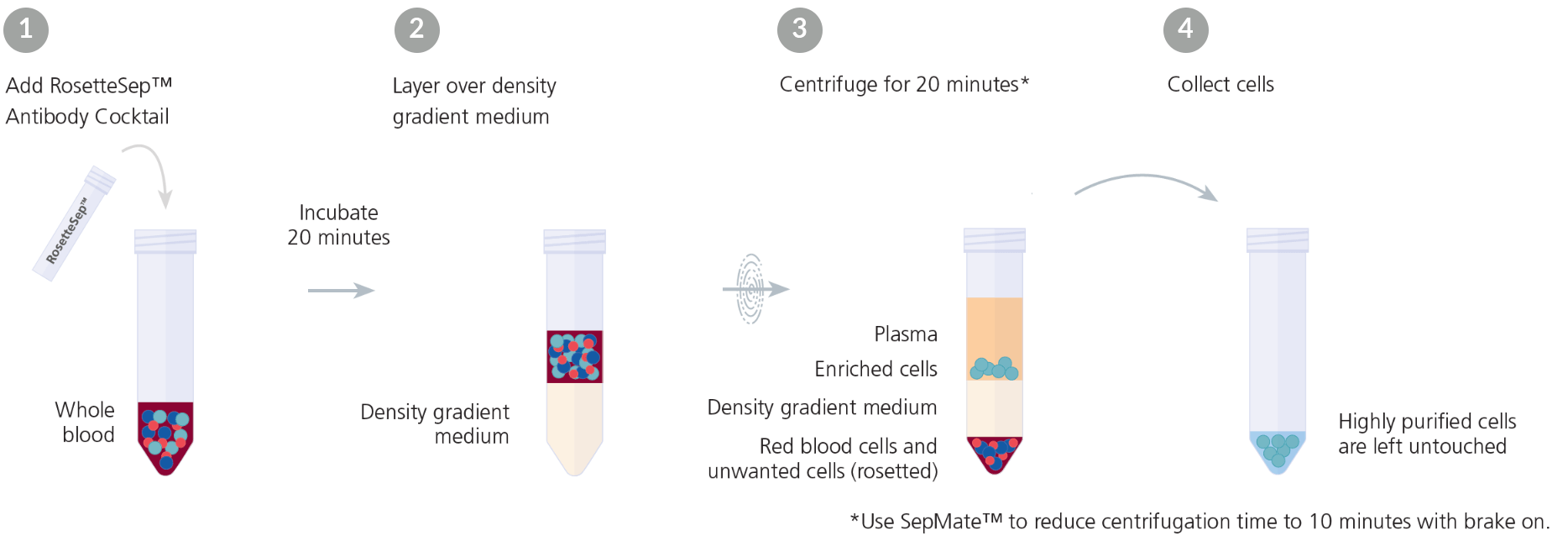
Figure 1. Typical RosetteSep™ Cell Separation Protocol
Sedimentation
Sedimentation works on the basis that gravity will cause larger and denser components to sediment faster than materials that are smaller and less dense. The largest and densest components in a sample can be pelleted through an initial low-force centrifugation due to their high rate of sedimentation. The supernatant can then be spun again. Through successive centrifugations, components with an increasingly lower rate of sedimentation can be isolated. Leukocytes are commonly separated from erythrocytes through dextran sedimentation. HetaSep™ is an example of an erythrocyte aggregation agent that is used to separate nucleated cells from red blood cells (RBCs) in whole blood.
Sedimentation is inexpensive but generally results in lower purity than other methods.
The unique adhesion profiles of different cell types can be used to separate target cells from heterogeneous populations. By choosing suitable growth factors and cell culture plates to selectively favor or inhibit adhesion, adherent cells can be separated from cells in suspension.
Macrophages are inherently adherent and often isolated from peripheral blood and bone marrow by adhesion. Mononuclear cells can be cultured with serum and a differentiation cocktail, promoting the formation of an adherent monolayer of macrophages. After removing the supernatant containing unwanted cells, the macrophages can be isolated.
Alternatively, cells that naturally grow in suspension or have lost anchorage dependency can be isolated by culturing the heterogeneous cell population in plates designed for ultra-low attachment. Without a surface to adhere to, adherent cells will fail to survive and the target cells will remain in suspension1.
Microfluidic Cell Separation
Microfluidics is an umbrella category of cell separation methods.2 Designed to manipulate fluids on a microscopic level to facilitate single-cell isolation, microfluidic technologies are frequently built onto microchips and are commonly known as "lab-on-a-chip" devices. These devices have several advantages, including the smaller volumes of samples and reagents required for use. Lab-on-a-chip devices are also portable, making them particularly useful as field-based diagnostic tools.
Microfluidic methods can be divided into active and passive systems. Active microfluidic systems involve external forces, whereas passive microfluidics make use of the cell’s density and mass in combination with gravity. These methods can also be classified by the presence or absence of cell labeling; although some methods involve labeling cells with antibodies, most methods are known for being label-free. There are several different microfluidic methods used for cell isolation, including:
- Acoustophoresis
- Aqueous two phase systems
- Biomimetic microfluidics
- Cell affinity chromatography
- Deterministic lateral displacement
- Electrophoretic sorting
- Field flow fractionation
- Gravity and sedimentation
- Magnetophoresis
- Microfiltration
- Optical sorting
Other Cell Separation Techniques
This section summarizes other, less commonly used, cell separation methods.
Aptamer Technology
Aptamers are single-stranded RNA or DNA oligonucleotides that form structures that can bind to highly specific targets. Through systematic evolution of ligands by exponential enrichment (SELEX) technology, aptamers can be screened and synthesized to target any cell type. These aptamers have high affinity and specificity toward their targets, and can be labeled with fluorochromes or magnetic particles to facilitate cell separation. The main advantage of aptamers is that they lack immunogenicity.
Fluorophore-labeled aptamers have been used to sort mesenchymal stem cells3 from bone marrow and RNA aptamers have been used to isolate mouse embryonic stem cells2.
Buoyancy-Activated Cell Sorting
Buoyancy-Activated cell sorting is a cell separation technique that utilises glass microbubbles labeled with antibodies specific to the target cells. When mixed into the sample, the microbubbles bind to the target cells. Due to the augmented buoyancy force, the microbubbles float to the surface, separating the target cells.
Complement Depletion
The complement depletion method takes advantage of the proteolytic cascade initiated by the complement system of the immune system. The complement system consists of plasma proteins that can be activated by pathogens or antibodies. Once activated, the plasma proteins induce the formation of a membrane-attack complex on a cell, resulting in cell lysis. With specific monoclonal antibodies, any cell population can be targeted and lysed through the complement cascade.
Laser Capture Microdissection
Laser capture microdissection (LCM) is a technique that uses a narrow laser beam to cleave target cells or areas from mostly solid tissue samples. Through microscopic visualization, LCM can isolate cell populations from heterogeneous mixtures using cell morphology or specific histological and immunological staining. LCM is particularly useful when working with small sample sizes.
Immunoguided Laser Capture Microdissection
Immunoguided laser capture microdissection combines immunostaining with laser capture microdissection (see above). This allows immunophenotypes to be used, in addition to morphology and tissue location, to identify and isolate target cells from the tissue sample. This technique employs immunohistochemistry or immunofluorescence to guide the dissection process for isolating cells expressing a specific molecular marker, and is particularly useful when histological stains do not recognize certain cell populations.
Limiting Dilution
Limiting dilution involves isolating single cells through the dilution of a cell suspension. This technique can be carried out with standard pipetting tools and is commonly used to produce monoclonal cell cultures and single cell cultures for single-cell analysis4.
Micromanipulation
Micromanipulation, a form of manual cell picking, is a cell isolation technique involving the use of an inverted microscope and ultra-thin glass capillaries connected to an aspiration and release unit. The system moves through motorized mechanical stages, allowing the operator to carefully select a specific cell and apply suction via micropipette to aspirate and isolate the cell.
Our Cell Separation Technologies
We have developed efficient technologies to support a few of the most prevalent cell separation methods, including immunomagnetic and immunodensity cell isolation. Learn more about our technologies:
EasySep™ Immunomagnetic Cell Separation
Isolate cells in as little as 8 minutes with a simple pour.
RosetteSep™ Immunodensity Cell Separation
Isolate human cells directly from whole blood during density gradient centrifugation.
On-Demand Course on Human Immune Cell Isolation
Learn how to effectively process human samples and isolate highly pure target cells with EasySep™ in this free self-paced course.
Cell Separation Guide E-Book
Download a practical guide, complete with step-by-step protocols, to learn everything you need to know about common cell isolation techniques.
Explore these resources
References
- Colter DC et al. (2000) Rapid expansion of recycling stem cells in cultures of plastic-adherent cells from human bone marrow. Proc Natl Acad Sci U S A 97(7):3213-8.
- Dalili A et al. (2018) A review of sorting, separation and isolation of cells and microbeads for biomedical applications: microfluidic approaches. Analyst 144(1):87-113.
- Guo KT et al. (2009) A new technique for the isolation and surface immobilization of mesenchymal stem cells from whole bone marrow using high‐specific DNA aptamers. Stem Cells 24(10):2220-31.
- Gross A et al. (2015) Technologies for single cell isolation. Int J Mol Sci 16(8): 16897-919.