iPSCdirect™
Single-use, maintenance-free human induced pluripotent stem cells, frozen
Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration
Overview
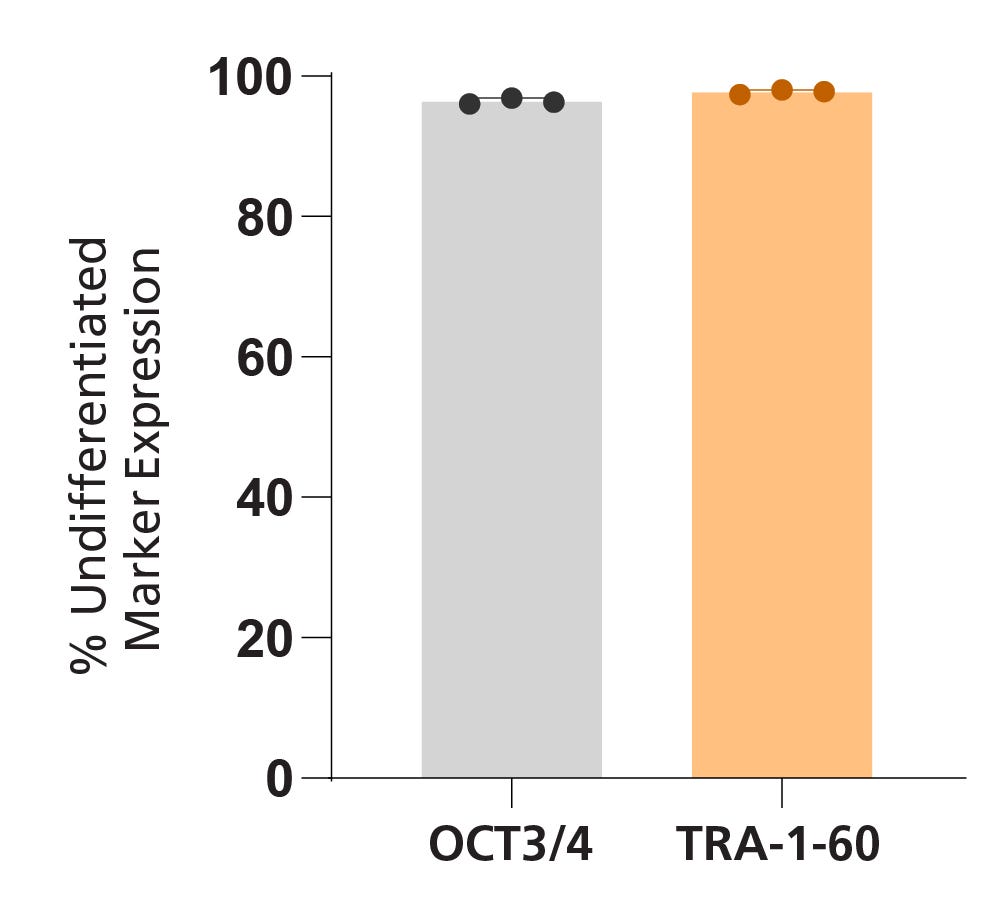
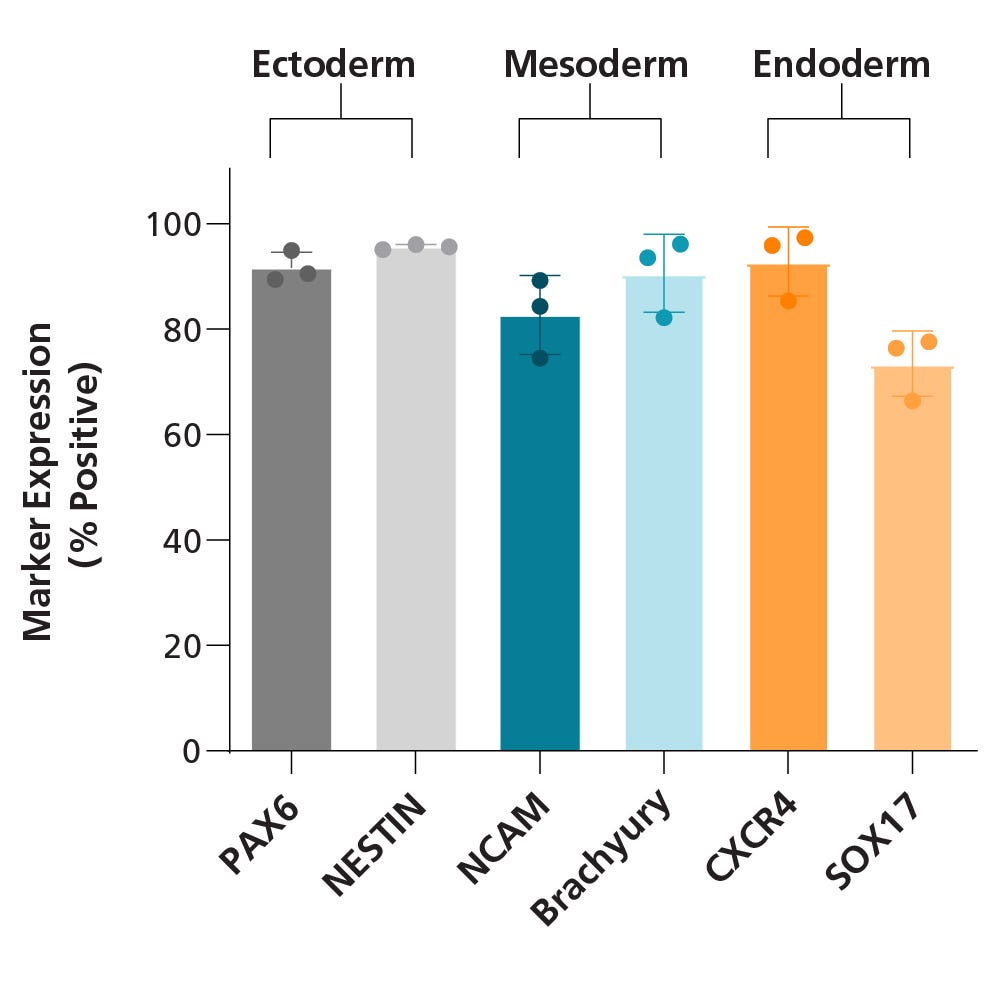
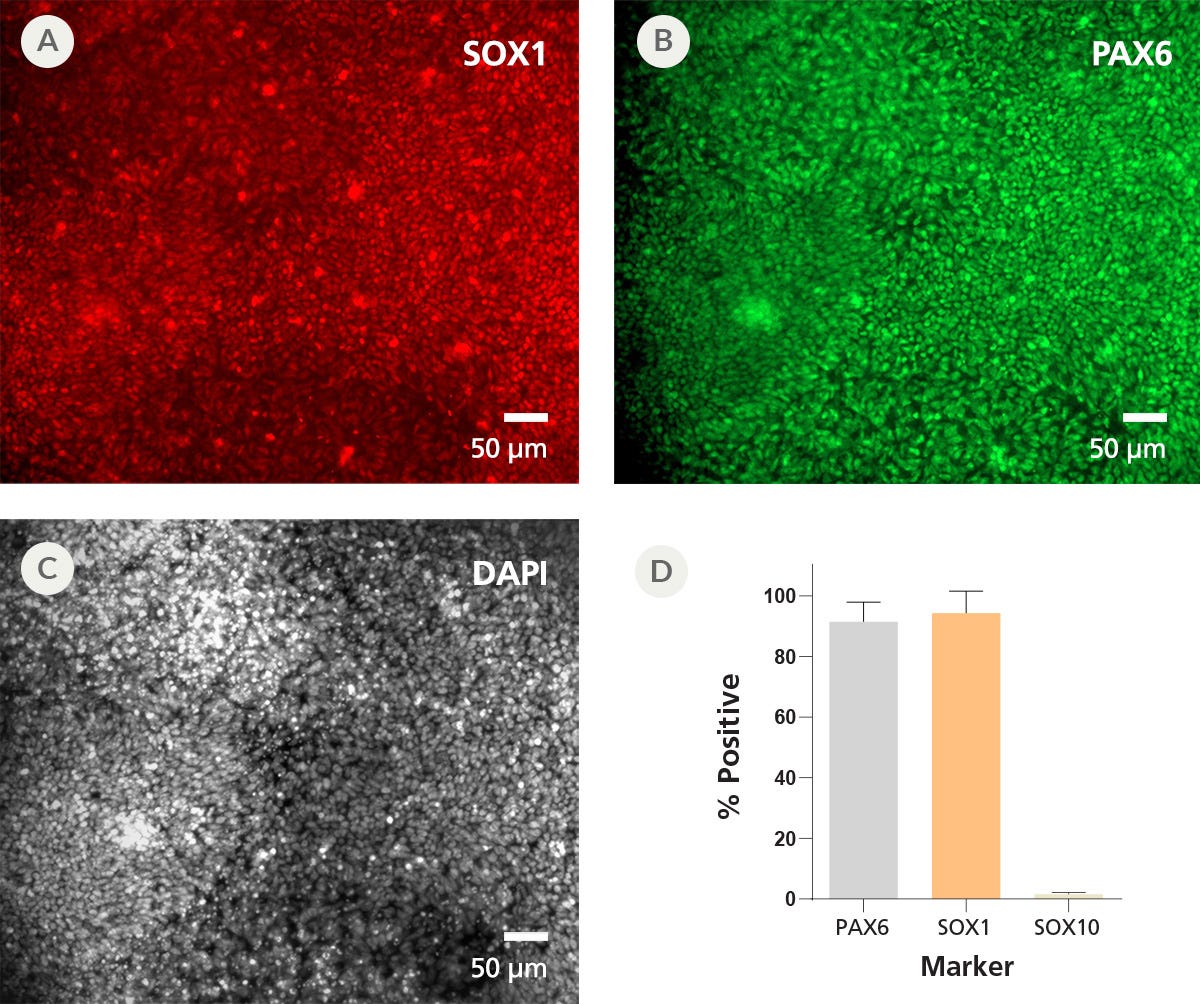
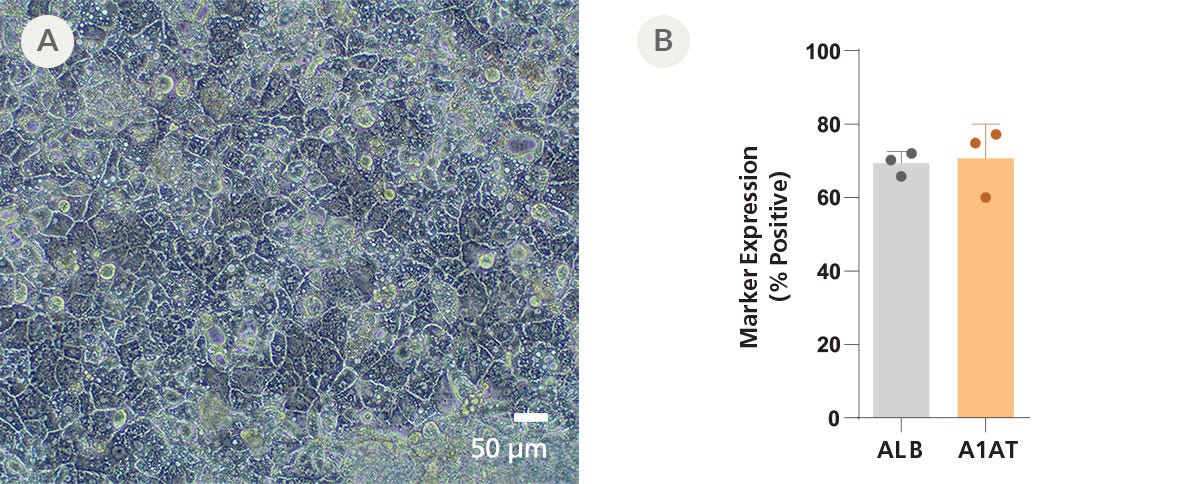
Data Figures
Protocols and Documentation
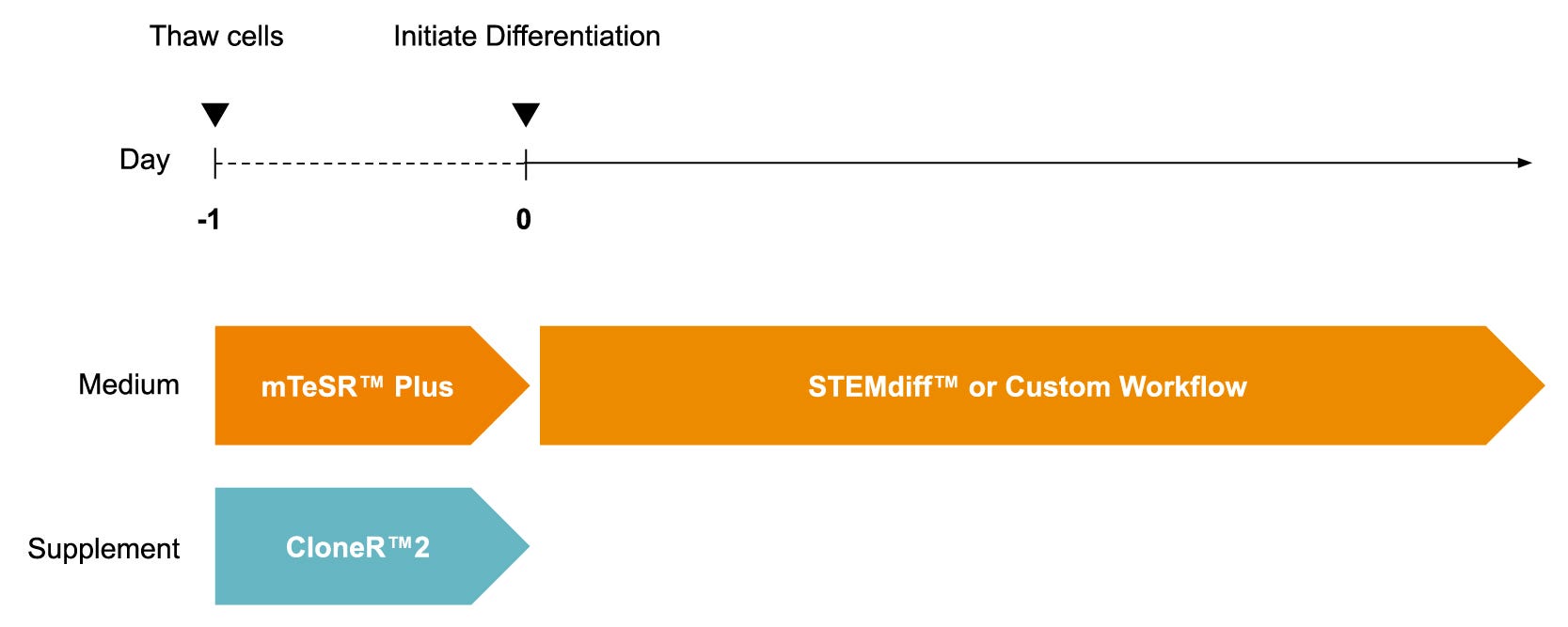
Find supporting information and directions for use in the Product Information Sheet or explore additional protocols below.
Applications
This product is designed for use in the following research area(s) as part of the highlighted workflow stage(s). Explore these workflows to learn more about the other products we offer to support each research area.
Resources and Publications
Educational Materials (10)
LIMITED USE LICENSEiPSCdirect™ is a single-use only product. Long-term maintenance or culture of iPSCdirect™ is not permitted. To maintain or expand iPSCdirect™ in the undifferentiated state, end users are required to sign the Standard License Agreement for iPSCs and an annual license fee will apply. Please contact iPSCrequests@stemcell.com for more information.These iPSCs and their modifications (including but not limited to derivatives or differentiated progeny) shall not be used or administered in (1) human subjects for human clinical use; (2) animals for veterinary use for therapeutic, diagnostic, or prophylactic purposes or (3) any subject in relation to, without limiting the generality of the foregoing, clinical applications, cell therapy, transplantation, and/or regenerative medicines, without limiting the generality of the foregoing. These iPSCs and their modifications (including but not limited to derivatives or differentiated progeny) may not be used for monetization or commercialization purposes, including without limitation, used to, or with the goal to, perform services or supply products or rights, including in the manufacture of cellular therapies or other therapeutics, for monetary gain or the generation of royalties, revenues, sales or other valuable consideration. For clarity, these iPSCs and their modifications (including but not limited to derivatives or differentiated progeny) may not be used for screening compounds, antibodies, proteins or peptides, except for the purposes of target discoveryF, target validation, or assay development, provided such activities and the results of such activities are not further used for monetization or commercialization purposes. It may be possible to obtain a further license for the prohibited uses referred to in this Limited Use License. Please contact iPSCrequests@stemcell.com for more details.
PRODUCTS ARE FOR RESEARCH USE ONLY AND NOT INTENDED FOR HUMAN OR ANIMAL DIAGNOSTIC OR THERAPEUTIC USES UNLESS OTHERWISE STATED.