Culture and Analysis of Hematopoietic Progenitor Cells in CFU Assays
Use these tips to better understand how the right medium can make the analysis of your hematopoietic CFU assays clearer
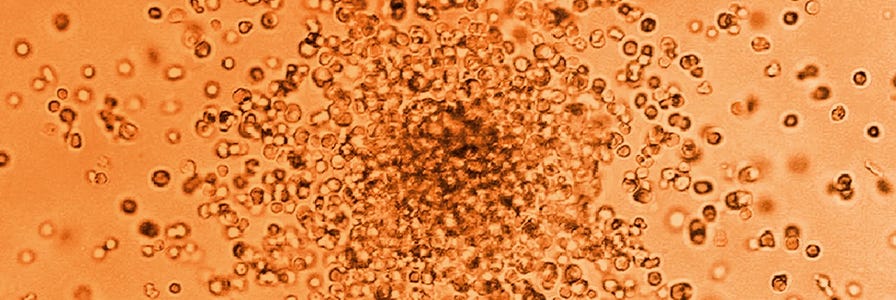
The colony-forming unit (CFU) assay is one of the most widely used assays for hematopoietic stem and progenitor cells (HSPCs). CFU assays allow measurement of the proliferation and differentiation ability of individual cells within a sample. The potential of these cells is measured by the observation of the colonies (consisting of more differentiated cells) produced by each input progenitor cell. About 14 days of culture is sufficient to allow colonies to grow to a size which allows accurate counting and identification, though shorter periods may be used in certain situations.
Previously, we covered topics related to the CFU assay such as determining the plating density appropriate for hematopoietic cell samples enriched with HetaSep™, and the importance of ensuring proper hydration during culture. The next step for correctly using the CFU assay is to have proper identification and counting methods to analyze the colonies that have grown.
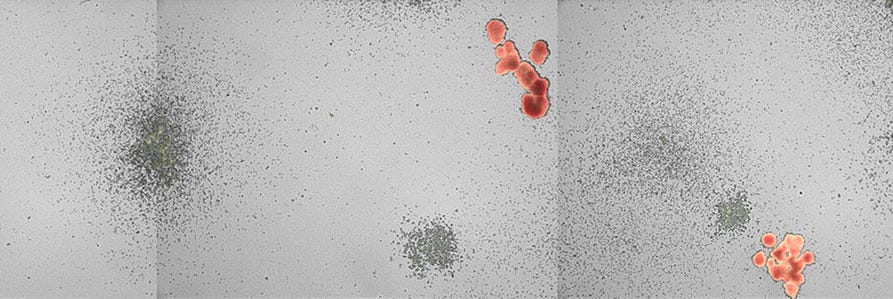
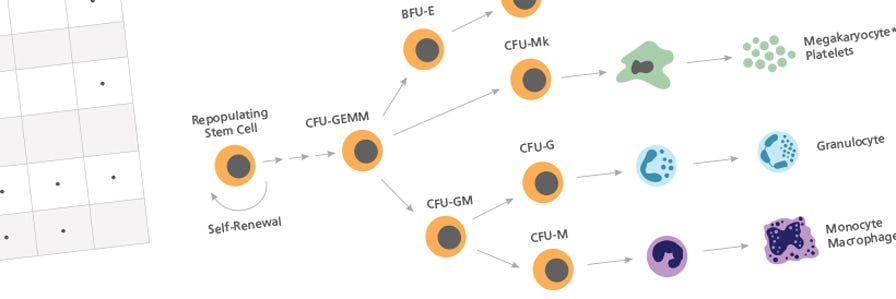
The numbers and types of colonies grown in culture are dependent both on the cell population plated and on the MethoCult™ medium used. Complete MethoCult™ media (e.g. MethoCult™ H4434 Classic) contain cytokines, including erythropoietin (EPO), that support the formation of granulocyte, monocyte, erythroid and mixed colonies, which contain all three lineages in addition to megakaryocytes. These colonies can be identified on the basis of the morphology of the cells and, in the case of erythroid and mixed colonies, the red colour observed in cells expressing hemoglobin.
MethoCult™ Media for Hematopoietic CFU Assays
Use the “Gold Standard” media for the in vitro detection and quantification of hematopoietic progenitor cells in the CFU assay.
It is not always easy to accurately identify the different colony types present in cultures, particular for those who are new to this assay, or are using atypical cell sources or samples. Training by experienced users is often an essential part of becoming proficient at counting and classifying colonies in the CFU assay.
One way to simplify the analysis of CFU assays while still obtaining information about lineage-specific progenitor cell growth is to perform assays in lineage-specific MethoCult™ media (see Tables 1 and 2) in which all colonies are derived from progenitor cell subtype, e.g., only CFU-GM or BFU-E. Colony counting and classification can also be done automatically with the STEMvision™ instrument for imaging and analysis of human and mouse CFU assays. Analysis software comes in the form of algorithms designed specifically for particular MethoCult™ medium formulations (Tables 1 and 2). An important advantage of STEMvision™ is that it takes away the subjectivity of manual counting and allows for standardization in the analysis of CFU assays within and between laboratories. STEMvision™ is also useful for applications in which manual colony identification and counting is prefered, as the software takes high resolution images of each culture well. These images can be used as a basis for manual counting at any time, with the added benefit of creating permanent records of cultures which may be revisited or reanalyzed at a later time.
Training: Introduction to STEMvision™ for Automated CFU Scoring
Learn how to standardize and automate the scoring of the CFU assay using STEMvision™ in this self-paced digital course. You’ll gain access to lectures, step-by-step protocol videos, and a library of curated resources.
CFU Assays with Human Samples
Human CFU assays may be performed in medium which supports the growth of multiple progenitor cell lineages, or in lineage-specific medium which facilitates the use of total manual counts. Automated counts with STEMvision™ instrument provide counts of colonies and classification into progenitor cell subtypes for human samples.
Table 1. Medium for Human Progenitor Cell Subsets
- Erythroid (BFU-E, CFU-E with BM)
- Myeloid (CFU-G/M/GM)
- Mixed (CFU-GEMM)
* STEMvision™ algorithms are available for counting cord blood-**, bone marrow- or peripheral blood-derived colonies.
**MethoCult™ Express medium is also available for performing total colony counts of cord blood samples at 7 days, with the STEMvision™ Human Cord Blood 7-Day algorithm for analysis.
Identification of Colonies Derived from Human Hematopoietic Progenitors
Get a quick reference of representative colony images and tips for identifying progenitor subtypes in CFU assays.
CFU Assays with Mouse Samples
Mouse CFU assays may be performed in general medium to promote the growth of granulocyte/macrophage and erythroid colonies in the same culture conditions, or in lineage-specific media that support granulocyte/macrophage or erythroid colony formation respectively. STEMvision™ algorithms are available for generating total counts of colonies in each of these media, providing information about multilineage, myeloid and erythroid colony potential, if assays are performed in parallel using the same cell sample in the specific media formulations indicated below.
Table 2. Medium for Mouse Progenitor Cell Subsets
- Myeloid (CFU-G/M/GM)
- Erythroid (BFU-E)
- Mixed (CFU-GEMM)
*STEMvision™ algorithms are available for use with CFU assays of mouse bone marrow only.
Identification of Colonies Derived from Mouse Hematopoietic Progenitors
Get a quick reference of representative colony images and tips for identifying progenitor subtypes in CFU assays.
For more information about performing and analyzing CFU assays with MethoCult™ and STEMvision™ please contact techsupport@stemcell.com.
Explore additional resources and support for setting up, optimizing, and standardizing the CFU assay.
Scientific Resources
Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration