Culturing Leukemic Stem & Progenitor Cells with StemSpan™ Medium
Protocol Describing How to Culture Chronic Myeloid Leukemia Cells with StemSpan™ Serum-Free Expansion Medium II (SFEM II)
- Document # 27105
- Version 2.0.1
- Nov 2019
This Technical Bulletin describes the use of serum-free StemSpan™ medium and supplements for the culture of hematopoietic stem and progenitor cells (HSPCs) isolated from patients with chronic myeloid leukemia (CML) or acute myeloid leukemia (AML). The protocols described below have been adapted from those originally developed for the isolation and culture of normal HSPCs, and enable the efficient in vitro expansion of CD34+ cells isolated from CML and AML stem and progenitor cells, even when initial cell numbers are low or cell quality is poor.
Applications of Leukemic Cell Culture
- Research into the development of new therapies for CML or AML
- Study of disease progression and clonal evolution of leukemic stem and progenitor cells
- Elucidate mechanism of resistance and relapse of leukemic stem and progenitor cells to current therapies
- Identify biomarkers and develop assays to predict patient response to novel therapeutics
Background
Leukemias are a group of disorders of the blood and bone marrow and are characterized by malignant transformation and clonal expansion of HSPCs.
CML is caused by a reciprocal translocation between chromosomes 9 and 22, yielding the Philadelphia (Ph) chromosome (Figure 1). The fusion gene encodes the BCR-ABL oncoprotein, a constitutively active tyrosine kinase that drives pathogenesis by perturbing multiple signaling cascades including the RAS/MAPK, PI3K/AKT, and JAK2/STAT5 pathways. Deregulation of these pathways leads to increased cell proliferation and reduced apoptosis, resulting in a massive accumulation of myeloid cells (predominantly granulocytes) in circulation.
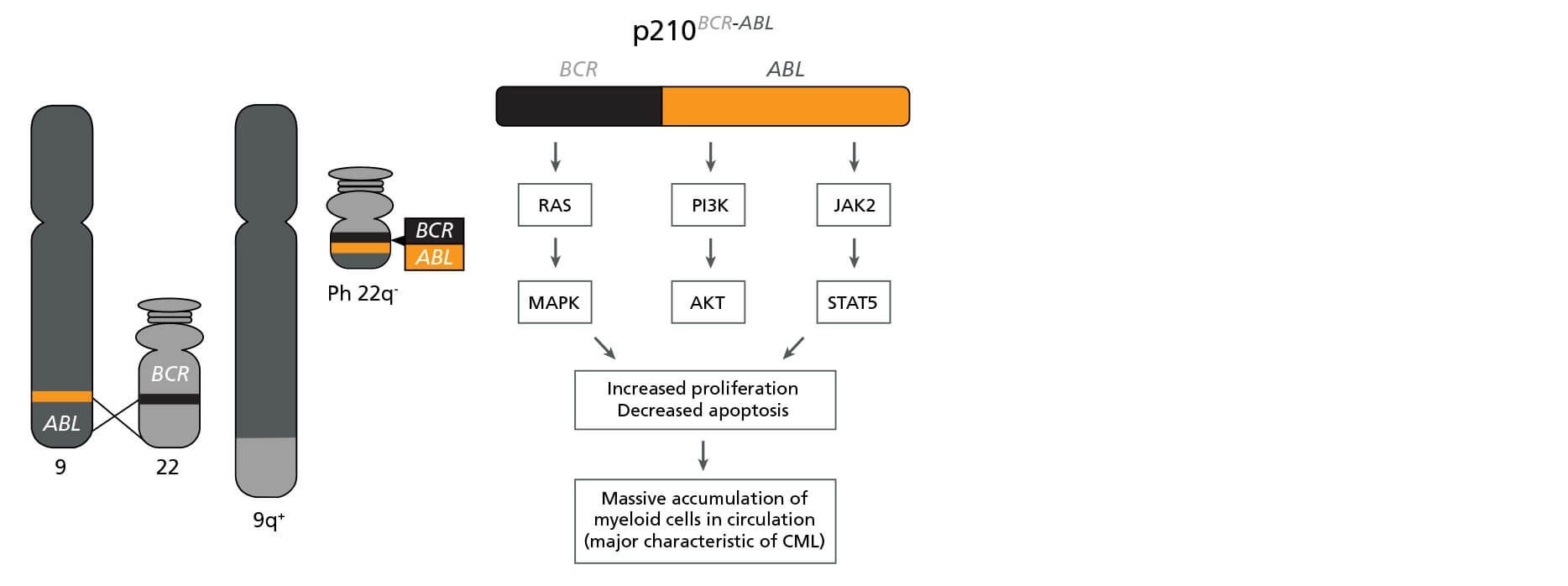
Figure 1. The BCR-ABL Fusion Gene
Schematic representation of the formation of the Philadelphia (Ph) chromosome encoding BCR-ABL and its biological effects. Adapted from Lydon, 2009.6
Due to the identification of this single specific mutation, CML has served as a model disease for other cancer types, allowing for the study of cancer evolution and development of molecular therapies that target the affected cells. The survival and prognosis of CML patients has improved in recent years thanks to the introduction of specific tyrosine kinase inhibitors, such as Imatinib and Dasatinib, that target the BCR-ABL oncoprotein and suppress the survival and proliferation of leukemic cells.
AML is much more heterogeneous than CML and is characterized by different chromosomal and genetic abnormalities with different risk profiles and different survival and relapse rates (Table 1).1 Furthermore, AML cells in individual patients often carry multiple genetic abnormalities, which play unique roles in disease initiation and progression.
Table 1. Common Genetic Abnormalities in AML.
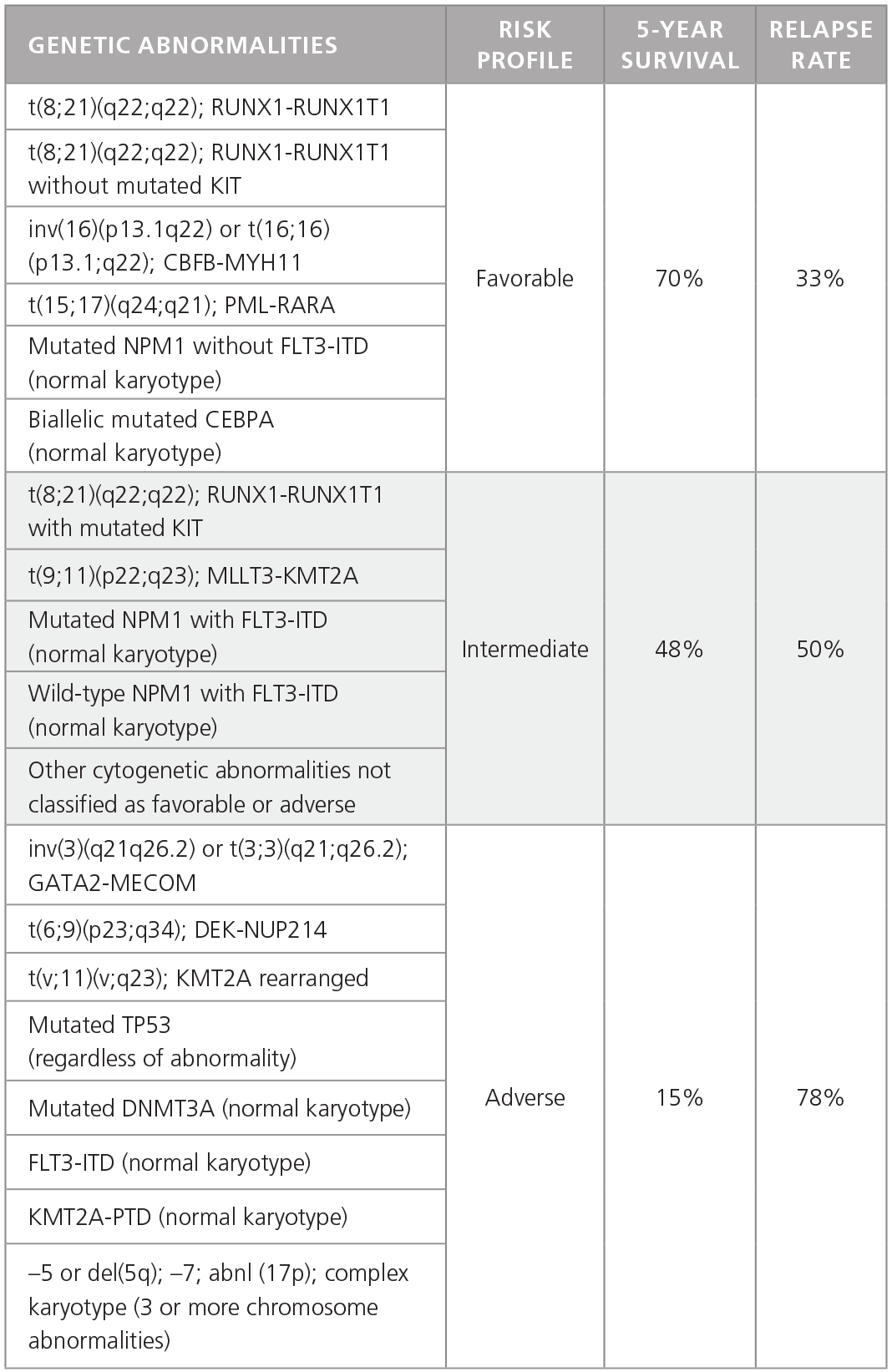
Adapted from Dohner H et al. 2015.1
The study of both CML and AML has been greatly facilitated by the use of in vitro culture systems similar to those used to study normal HSPCs. In vitro culture has been used to elucidate the basic biology of these diseases, while allowing conditions to screen compounds such as tyrosine kinase inhibitors or other candidate therapeutics on primary patient-derived cells.2-4 Studies on primary leukemic HSPCs are often impeded by the limited access to patient samples, which may be compounded by poor yield and viability of cryopreserved samples after thawing.
When culturing cells from valuable patient samples, it is important to choose a medium that is reliable, defined, and optimized for your cell type. The experiments presented in this Technical Bulletin were performed using the StemSpan™ Leukemic Cell Culture Kit (Catalog #09720), which contains StemSpan™ Serum-Free Expansion Medium II (SFEM II; Catalog #09605), StemSpan™ CD34+ Expansion Supplement (Catalog #02691), and the small molecule UM729 (Catalog #72332).5
Other media and supplements, e.g. StemSpan™ SFEM (Catalog #09600), StemSpan™ CC100 (Catalog #02690) or CC110 (Catalog #02697), and small molecules may also be used to generate similar culture conditions. The experiments in this Technical Bulletin were performed using the small molecule UM171, however similar results are expected when using UM729 prepared to a final concentration of 1μM.
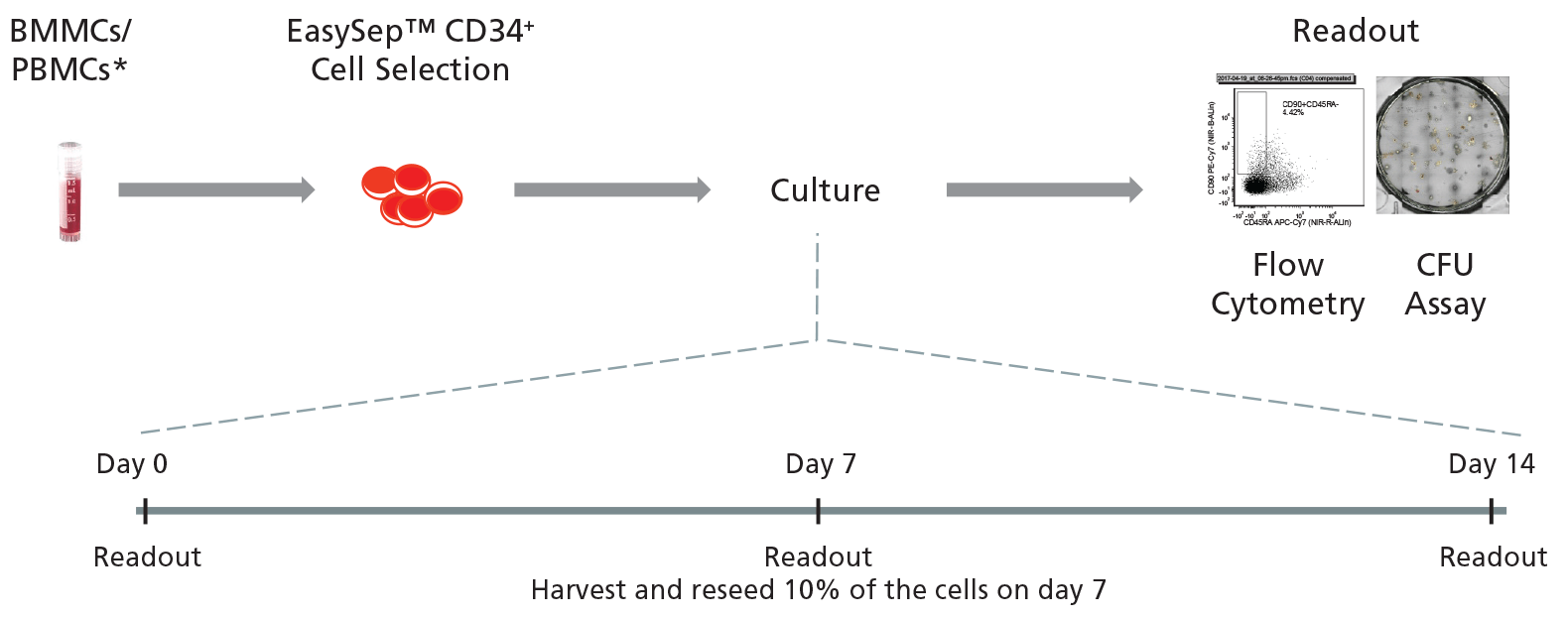
Figure 2. General Protocol for Culturing Leukemic CD34+ Cells.
CD34+ cells isolated from CML or AML samples may first be purified using EasySep™ Human Cord Blood CD34 Positive Selection Kit II (Catalog #17896) and cultured for 14 days in StemSpan™ SFEM II supplemented with StemSpan™ CD34+ Expansion Supplement with or without addition of UM729. Cells may be analyzed on days 0, 7, and 14 with colony-forming unit (CFU) assays using MethoCult™ H4435 Enriched (Catalog #04435) medium, and by immunophenotyping for cell surface markers commonly expressed on HSPCs.
*The experiments in this Technical Bulletin were performed using cryopreserved cells, however similar results are expected when using fresh samples. Additionally in place
of UM729, the small molecule UM171 was used to generate data in Figures 4 - 7. UM171 is no longer licensed for sale by STEMCELL Technologies, however similar results
are expected when using UM729 prepared to a final concentration of 1 μM (data not shown). Further titration may be necessary to optimize cell fold expansion in specific
conditions.
For more information including data comparing UM171 and UM729, see Fares et al. 2014.
Protocol
Isolate
- If starting with fresh peripheral blood mononuclear cells (PBMCs) or bone marrow mononuclear cells (BMMCs), proceed directly to step 2.
If starting with frozen PBMCs or BMMCs, thaw a 1 mL vial of sample in a 37°C water bath. When just thawed, add 0.5 mL of DNase I Solution (Catalog #07900), gently mix, and incubate at room temperature (15 - 25°C) for 5 minutes. Washing is not recommended prior to isolation of CD34+ cells as it may lower cell yield and viability. This thawing protocol increases cell yield and improves viability of the leukemic CD34+ cells. - Isolate CD34+ cells (from the DNase-treated thawed PBMCs/ BMMCs or fresh PBMCs from step 1), using EasySep™ Human Cord Blood CD34 Positive Selection Kit II (Catalog #17896). This kit was originally designed for use with cord blood, but has been extensively tested with CML and AML samples.
- Determine the number and percentage of viable cells using a hemocytometer with trypan blue or an automated cell counter. If enough cells are present, the percentage of CD34+ cells can also be measured using flow cytometry. The number and/or concentration of CD34+ cells can be determined by multiplying the total number of viable cells by the percentage of CD34+ cells in the sample.
Culture
-
Prepare 100 mL of leukemic cell culture medium by combining the following:
- 90 mL StemSpan™ SFEM II
- 10 mL StemSpan™ CD34+ Expansion Supplement
- UM729 to a final concentration of 1μM*
- Seed CD34+ cells at 1000 viable cells per well in 100 μL of the prepared leukemic cell culture medium in 96-well plates, or 1000-10,000 cells in 1 mL medium per well in 12-well plates.
- Incubate at 37°C for 7 days.**
- On day 7, harvest the cells, wash, and resuspend in 1 mL of Iscove's MDM without any cytokines or supplements added.
- Reseed 10% of the cell suspension into fresh leukemic cell culture medium, prepared as described in step 4,and incubate at 37°C for an additional 7 days.
- On day 14, harvest cells, wash, and resuspend as in step 7.
*If using UM171, prepare to a final concentration of 175 nM.
**Culture length can be modified based on the goals and requirements of the experiment. If culturing cells for >7
days, it is recommended to follow steps 7 and 8 at day 7, and as appropriate every additional 7 days.
Analyze
-
- Determine the number and phenotype of cells expressing HSPC markers CD45, CD34, CD45RA, and CD90 on day 0 (if enough cells are available after CD34+ cell isolation), and on day 7 and 14 after culture, using flow cytometry. Aldehyde dehydrogenase (ALDH) activity, which is high in primitive stem and progenitor cells, can also be determined using ALDEFLUOR™ (Catalog #01700) in parallel with HSPC marker staining.
- Measure the frequency and total number of hematopoietic progenitor cells on day 0, and on day 7 and 14 after culture, by performing colony assays with MethoCult™ H4435 Enriched Medium (Catalog #04435) (refer to the StemSpan™ Leukemic Cell Culture Kit Product Information Sheet (PIS; Document #DX23137). Count colonies after 14 days.
Optional: Colonies may be plucked from CML samples at day 14 for further analysis, including cDNA preparation and qPCR to determine BCR-ABL status. See Table 2 for primers. For AML samples, sequencing or qPCR may be performed to identify the presence of fusion genes or other mutations.
Results
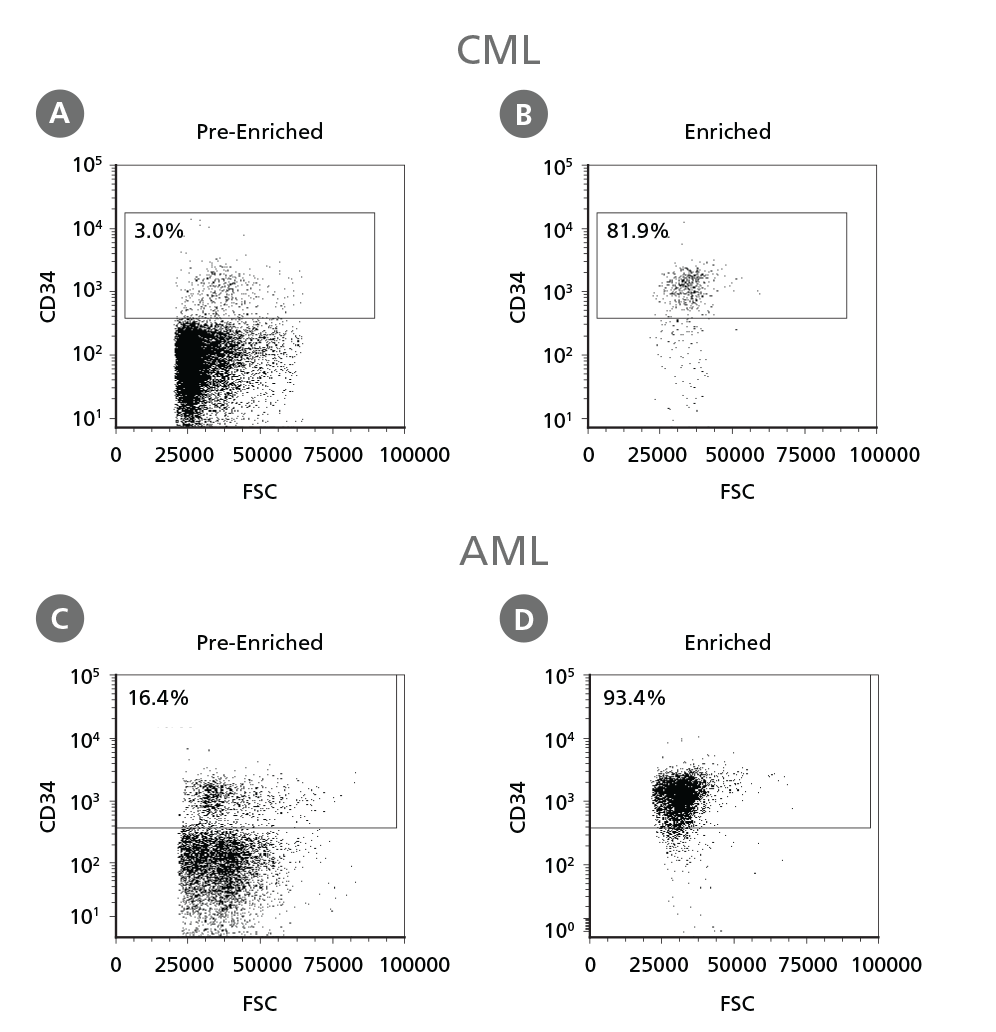
Figure 3. Isolation of Leukemic CD34+ Cells.
Cryopreserved CML or AML PBMCs and BMMCs were thawed and treated as described in Step 1 of the protocol. CD34+ cells were isolated using EasySep™ Human Cord Blood CD34 Positive Selection Kit II. The percentage of CD34+ cells before (A, C) and after (B, D) CD34+ cell isolation was measured by flow cytometry. Dead cells were excluded by light scatter profile and viability staining. In this example the purity of CD34+ cells increased from 3% to 82% (CML) and from 16% to 93% (AML).
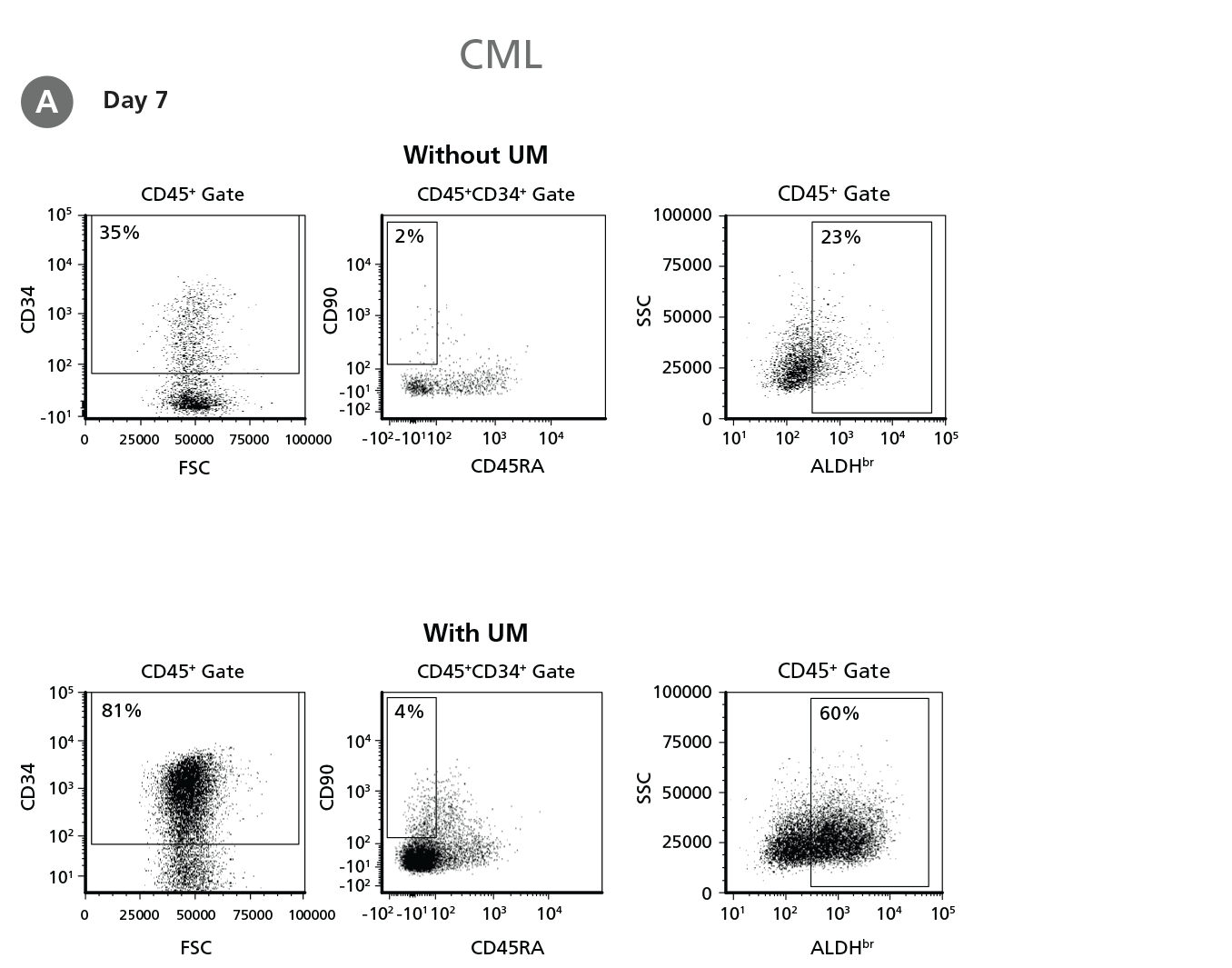
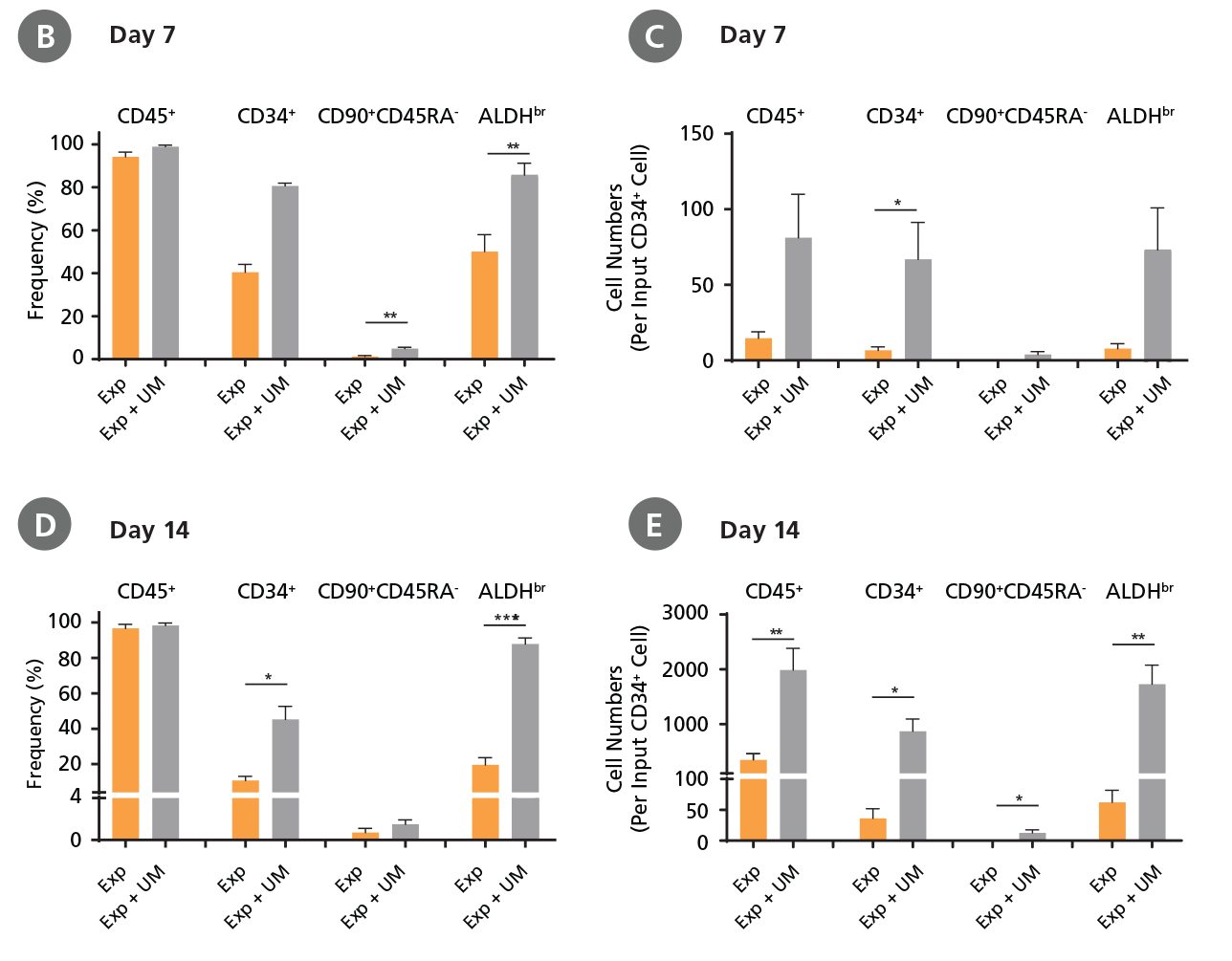
Figure 4. Expansion of CD34+ CML Cells.
CD34+ CML cells were cultured in StemSpan™ SFEM II containing CD34+ Expansion Supplement (Exp) without or with UM171 (UM) as described in the protocol. After 7 and 14 days, the cultured cells were stained with fluorescently labeled antibodies against CD45, CD34, CD90, CD45RA, and with ALDEFLUOR™ (Catalog #01700) to measure ALDH activity, and analyzed by flow cytometry. Sequential gates were used to determine the percentages of viable CD45+, CD45+CD34+, and CD45+CD34+CD90+CD45RA- cells (based on “Fluorescence Minus One" (FMO) controls), and ALDHbr cells (based on DEAB control). (A) Representative flow cytometry profiles at day 7 are shown. The (B,D) frequency and (C,E) cell numbers of these subsets per initial CD34+ cell on (B,C) day 7 and (D,E) day 14 are shown. StemSpan™ SFEM II supplemented with CD34+ Expansion Supplement supports the expansion of CML cells in culture. The addition of UM171 enhances expansion of all subsets shown (~10-fold expansion of CD34+ progenitor cells at day 7 and ~20-fold at day 14 compared to cultures without UM171). UM729 is expected to provide similar results when used at a final concentration of 1μM. Data shown are mean ± SEM (n = 6). P-values were calculated using a two-tailed paired Student's t-test (*P < 0.05; **p < 0.01; ***p < 0.0001). All six CML samples tested were able to expand in culture.
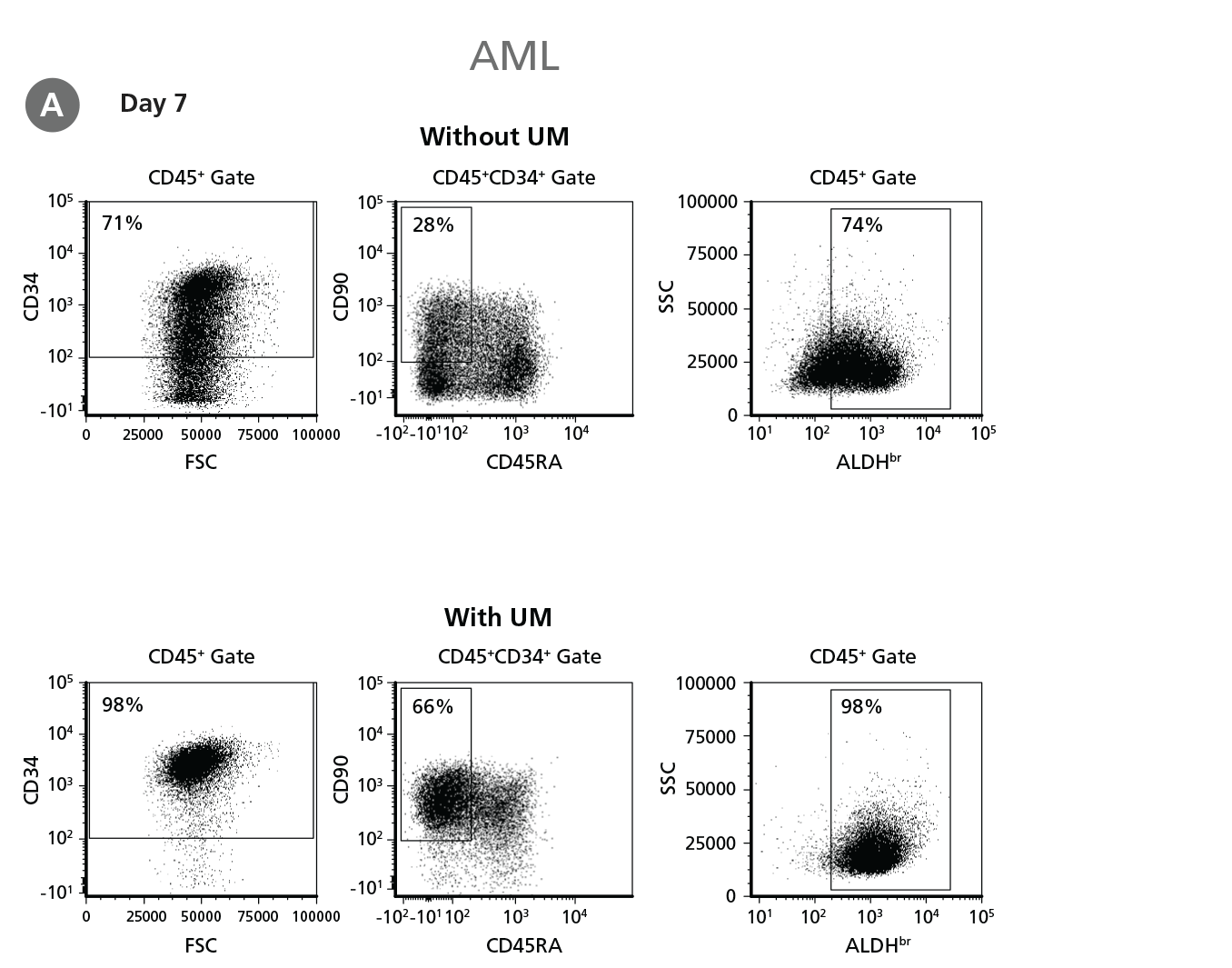
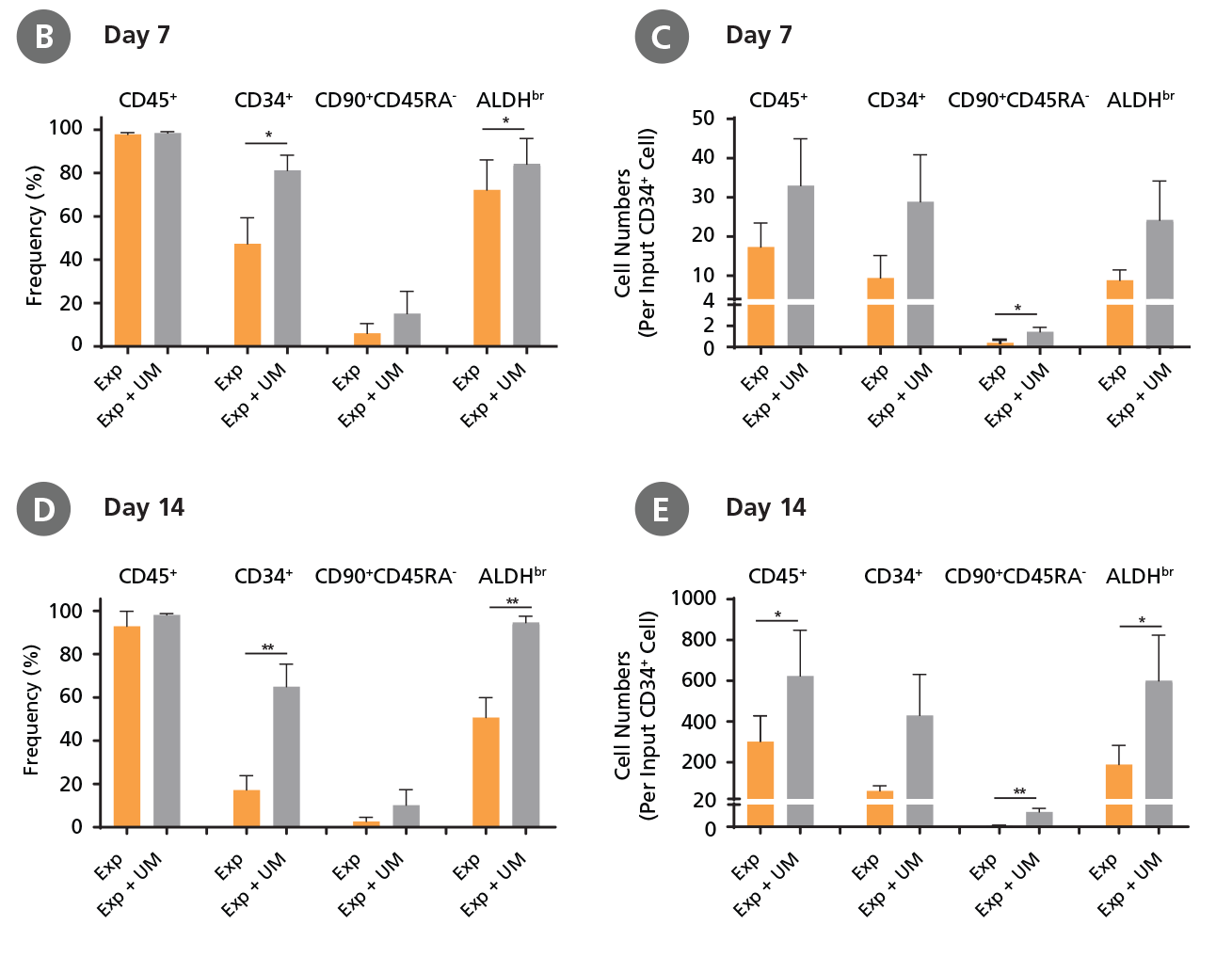
Figure 5. Expansion of CD34+ AML Cells.
CD34+ AML cells were cultured in StemSpan™ SFEM II containing CD34+ Expansion Supplement (Exp) alone, or with UM171 (UM) as described in the protocol. After 7 and 14 days, the cultured cells were stained with fluorescently labeled antibodies and with ALDEFLUOR™ as described in Figure 4. (A) Representative flow cytometry profiles at day 7 are shown. The (B, D) frequency and (C, E) cell numbers of these subsets per initial CD34+ cell on (B, C) day 7 and (D, E) day 14 are shown. SFEM II supplemented with CD34+ Expansion Supplement supports the expansion of AML cells in culture. The addition of UM171 further enhances expansion of all subsets shown (~3-fold expansion of all subsets at day 7 and ~7-fold at day 14 compared to cultures without UM171). UM729 is expected to provide similar results when used at a final concentration of 1μM. Data shown are mean ± SEM (n = 6). P-values were calculated using a two-tailed paired Student's t-test (*P < 0.05; **p < 0.01). Six out of ten AML samples tested were able to expand in culture.
When cultured with UM171, the frequencies of CD34+ cells, CD34+CD90+ CD45RA- cells, and ALDHbr cells in AML cultures at day 7 and day 14 (Figure 5) were similar to those of CML samples (Figure 4) (~80% CD34+ cells for both CML and AML samples at day 7); whereas the numbers of CD34+ cells, CD34+CD90+ CD45RA- cells, and ALDHbr cells in AML samples (Figure 5) were 2.5-fold lower than those of CML samples (Figure 4) (~70-fold expansion of CD34+ cells in CML samples vs ~30-fold expansion of CD34+ cells in AML samples at day 7).
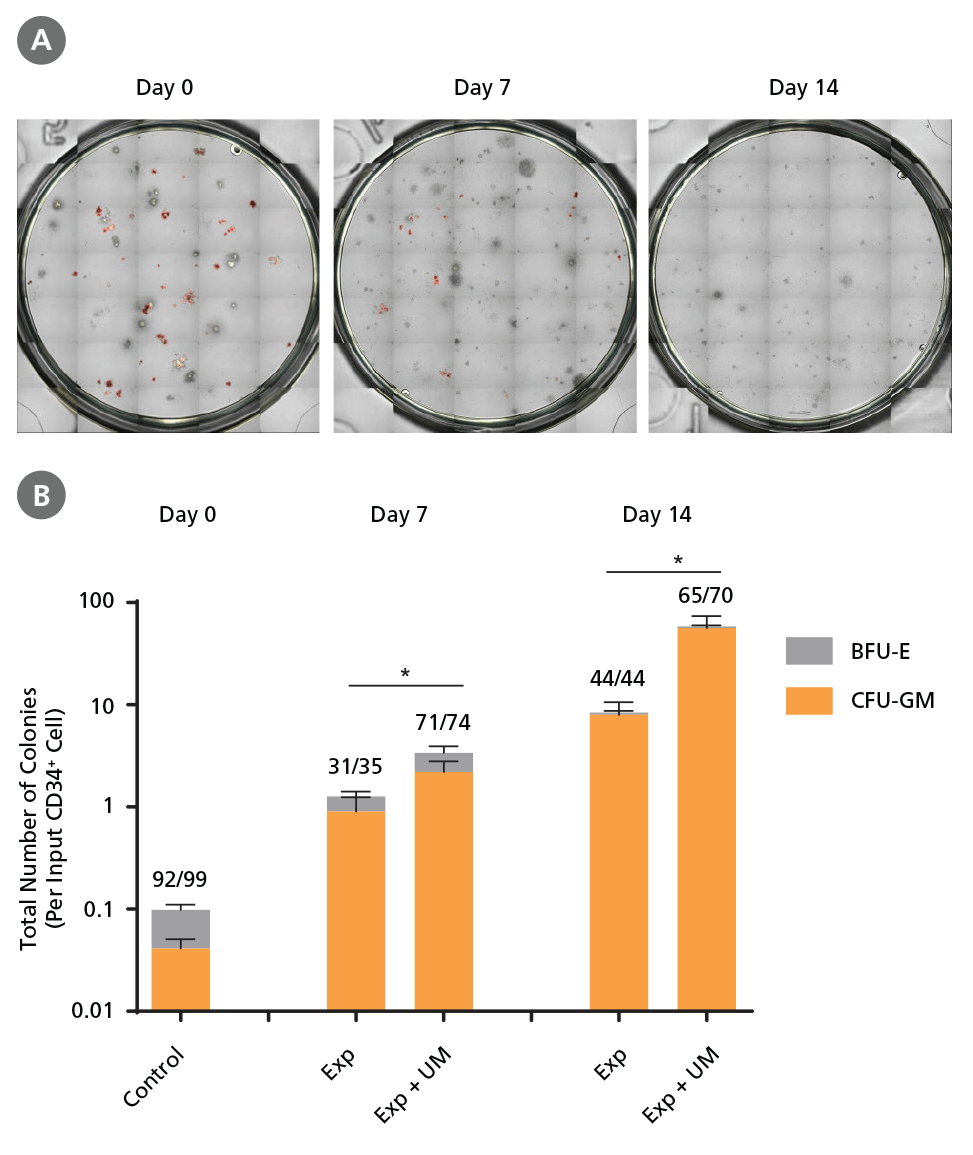
Figure 6. Colony-Forming Potential of CD34+ CML Cells is Maintained During Culture.
CML cells were assayed in colony assays using MethoCult™ H4435 Enriched medium directly after CD34+ cell isolation (day 0) or after 7 or 14 days of expansion without or with UM171 (UM; as described in Figure 4). After 14 days of culture in StemSpan™ SFEM II with SFEM II supplemented with CD34+ Expansion Supplement without or with UM171, colonies were (A) imaged with STEMvision™ and counted manually from digital images. (B) CFU output expressed as the total number of colonies per original input CD34+ cell. Numbers above each of the individual bars indicate the proportion of BCR-ABL positive colonies, measured by qRTPCR on individual plucked colonies across 6 different samples (8-12 colonies were plucked for each sample per condition, see Table 2 and Figure 8 for details on the primers and procedure). SFEM II supplemented with CD34+ Expansion Supplement (Exp) supports the expansion of colony-forming progenitor cells in culture. UM171 further promotes colony forming progenitor cell output (~3.5- fold expansion at day 7 and ~8-fold at day 14). UM729 is expected to provide similar results when used at a final concentration of 1μM. Single-colony qRT-PCR analysis revealed that colonies generated from samples on day 0, and colonies generated from cells expanded for 7 and 14 days, were predominantly BCR-ABL+ but also that normal BCR-ABL- progenitor cells were present at low frequencies. Data shown are mean ± SEM (n = 6). P-values were calculated using a two-tailed paired Student's t-test (*P < 0.05).
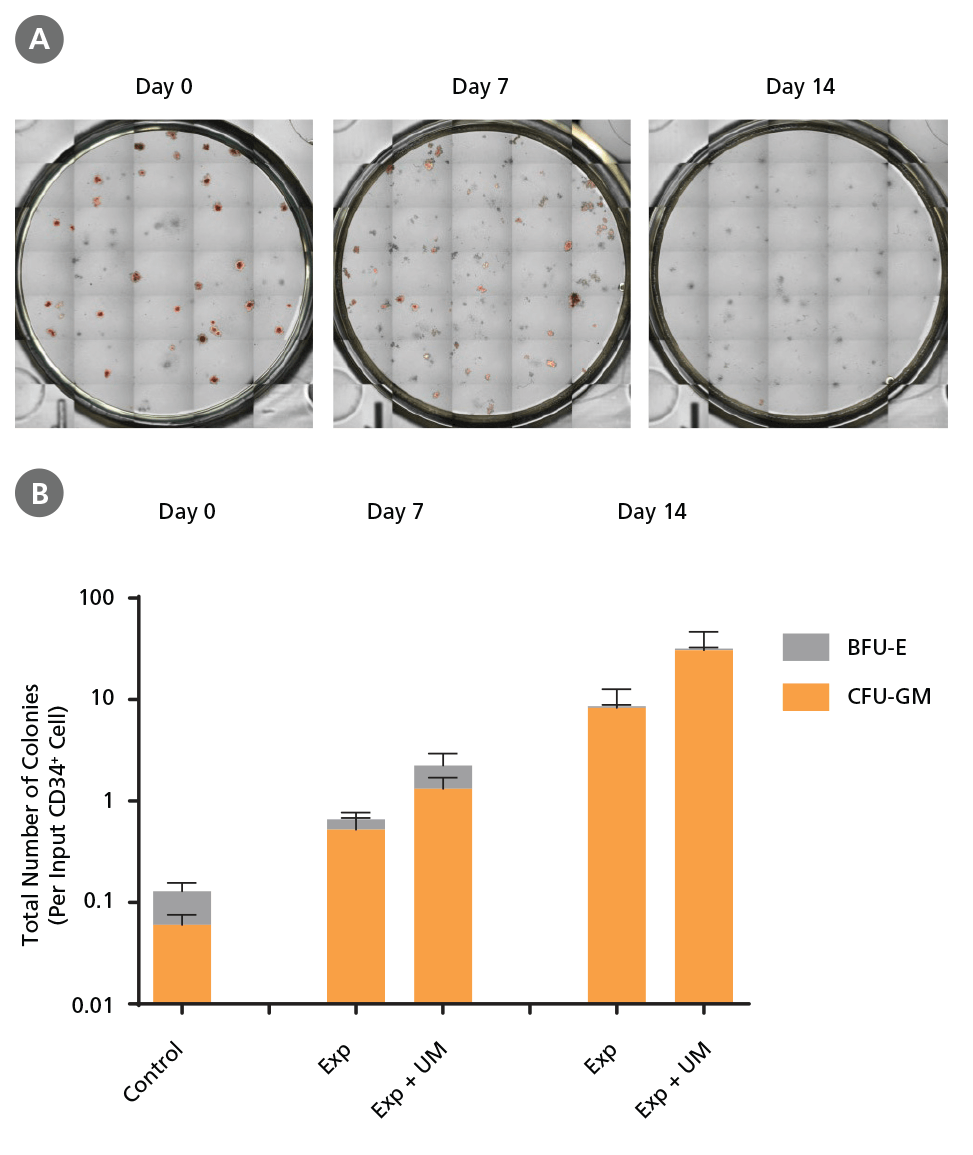
Figure 7. Colony-Forming Potential of CD34+ AML Cells is Maintained During Culture.
AML cells, after CD34+ cell isolation (day 0) or after 7 or 14 days of expansion without or with UM171 (UM; as described in Figure 4), were plated in colony assays with MethoCult™ H4435 Enriched medium. After 14 days of incubation, colonies were (A) imaged with STEMvision™ and counted manually from digital images. (B) CFU output expressed as the total number of colonies per original input CD34+ cell. SFEM II supplemented with CD34+ Expansion Supplement (Exp) supports the expansion of colony-forming progenitor cells in culture. Addition of UM171 further promotes colony-forming progenitor cell output (~3-fold expansion at day 7 and ~4-fold at day 14). UM729 is expected to provide similar results when used at a final concentration of 1μM. Data shown are mean ± SEM (n = 6). P-values were calculated using a two-tailed paired Student's t-test (*P < 0.05).
Table 2. Primer-Probe Sequences, Working Concentrations, and Primer Efficiencies.
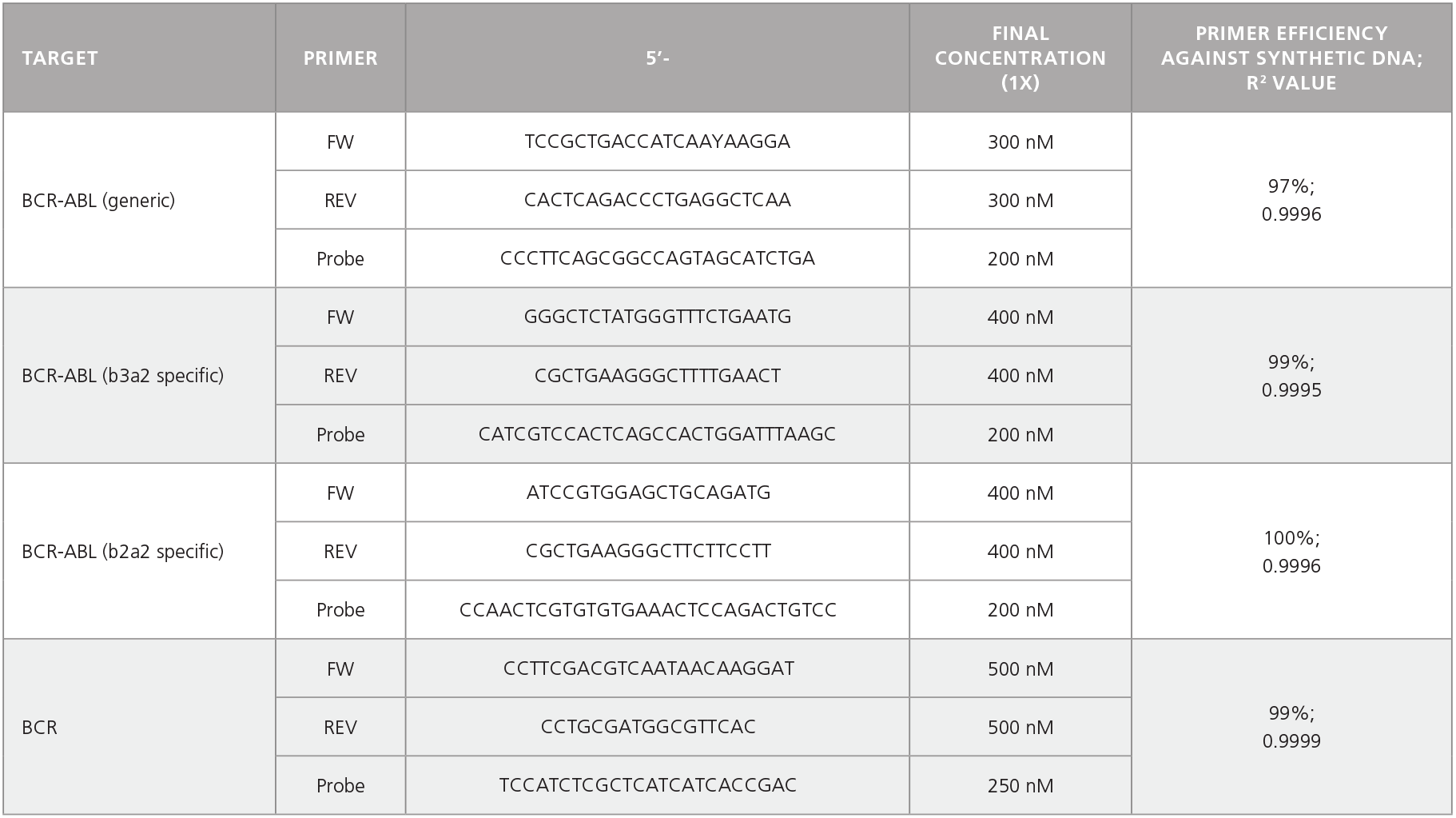
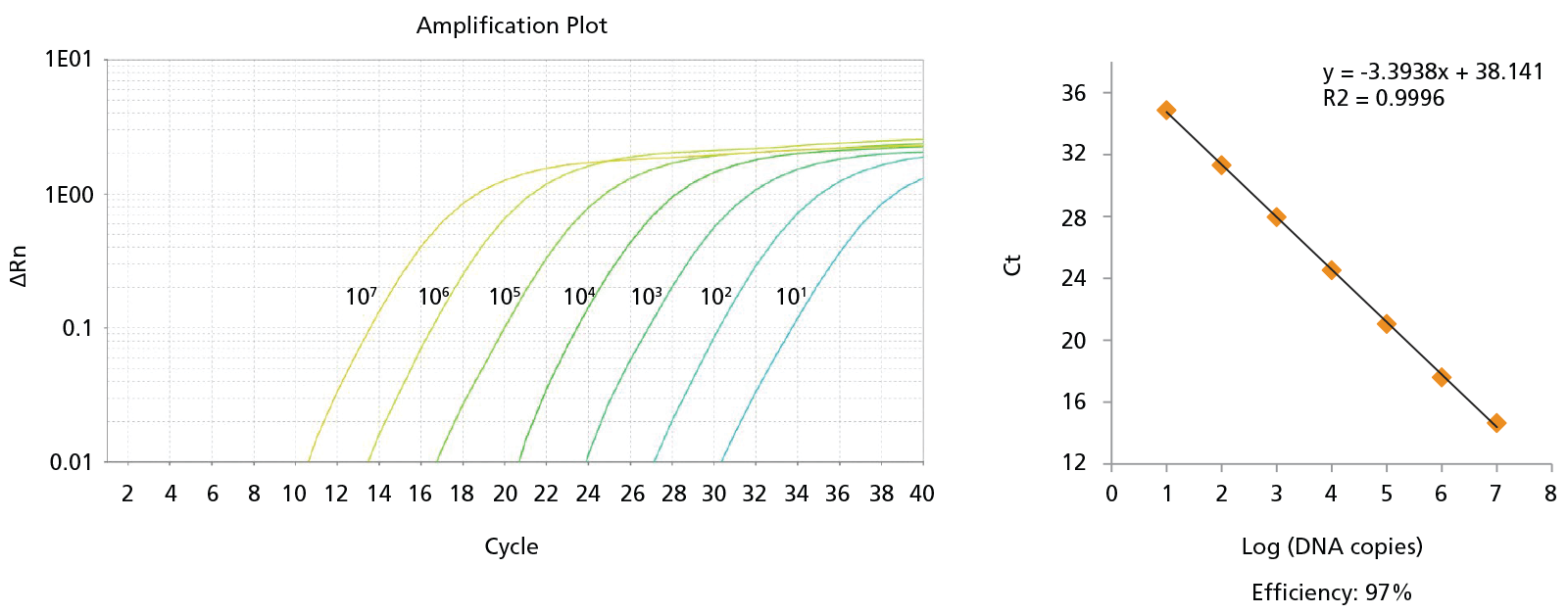
Figure 8. Probe-Based qPCR Assays Can Be Used to Identify Single BCR-ABL+ Colonies Generated in CFU Assays of CD34+ Cells Isolated from CML Samples.
Probe-based qPCR assays were utilized to identify BCR-ABL+ single colonies obtained from the non-expanded, 7-day-expanded, and 14-day-expanded CML cells as shown in Figure 6. All primer sets were able to detect as few as 10 copies of DNA with primer efficiencies > 90%. An amplification plot and a standard curve using BCR-ABL generic primer against b2a2 synthetic DNA fragments are shown. Numbers beside each amplification curve indicate number of DNA copies. Primer-probe sequences, working concentrations, and primer efficiencies are shown in Table 2.
Products
For Isolation and Culture
*Similar results can be obtained using UM171 at a final concentration of 1.75 μM in culture.
For Analysis
For Colony Assay and qPCR Analysis
References
- Döhner H et al. (2015) N Engl J Med 373(12): 1136–52.
- Pabst C et al. (2014) Nat Methods 11(4): 436–42.
- Liao Z et al. (2015) Mol Cancer Ther 14(8): 1777–93.
- Baccelli I et al. (2017) Blood Cancer J 7(2): e529.
- Fares I et al. (2014) Science (80- ) 345(6203): 1509–12.
- Lydon N. (2009) Nat Med 15(10): 1153–7.
Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration


