Weekend-Free Culture of Human Pluripotent Stem Cells in mTeSR™1 or TeSR™-E8™
.
- Document # 28071
- Version 1.0.1
- Dec 2016
Introduction
mTeSR™1 (Catalog #05850) is the most-published medium for the culture of human embryonic stem (ES) and induced pluripotent stem (iPS) cells in serum-free and feeder-free conditions. Established protocols using mTeSR™1 require daily medium replacement, with the accepted exception to allow one double-volume medium change followed by a single day without feeding each passage, causing researchers to routinely work at least one day on weekends.
To establish a completely weekend-free work schedule, we developed a protocol that involves routine passaging on Fridays and eliminates feeding on Saturdays and Sundays (when colonies are small). Development of this 7-day passaging schedule required optimization of the size and density of aggregates seeded. We validated this 7-day passaging protocol with cells cultured in mTeSR™1 and TeSR™-E8™ (Catalog #05940). Corning® Matrigel® hESC-qualified matrix (Matrigel®) or the defined protein Vitronectin XF™ (Catalog #07190; developed and manufactured by Primorigen Biosciences, Inc.) were used as matrices. Cultures maintained using this weekend-free protocol retained the same high-quality standard typically seen with TeSR™ media.
High Culture Quality is Maintained Without Weekend Feeding
We compared the ability of our newly developed weekend-free 7-day passaging protocol (Figure 1) to support long-term maintenance of human ES and iPS cells against a standard 7-day passaging protocol where medium was replaced 6 of 7 days with a double-volume feed on Saturdays and no feed on Sundays. In both cases, cells were routinely passaged on Fridays using the same protocol (See Protocol Tips & Tricks). Experiments were performed using H1 or H9 human ES cells, or WLS-1C or WLS-4D1 human iPS cells maintained in either mTeSR™1 or TeSR™-E8™ on Matrigel® unless otherwise indicated. For each experiment, duplicate wells were independently maintained in parallel for at least 10 passages.
Key Features:
- Passage once a week on Fridays
- No work required Saturdays or Sundays
- Protocol validated with mTeSR™1 and TeSR™-E8™
- Compatible with Matrigel® and Vitronectin XF™
Cultures maintained with the weekend-free 7-day passaging protocol retained the features of undifferentiated human ES and iPS cells. Markers of human pluripotent stem cells such as OCT-4, TRA-1-60 and SSEA-3 remained highly expressed for at least 10 passages (Figure 2). Cultures also exhibited consistently excellent colony morphology, as indicated by round colonies with well-defined edges, phase-bright colony centers, and minimal overt spontaneous differentiation (Figure 3). Cells within undifferentiated colonies were homogeneous and tightly packed with prominent nucleoli and a high nuclear-to-cytoplasmic ratio.
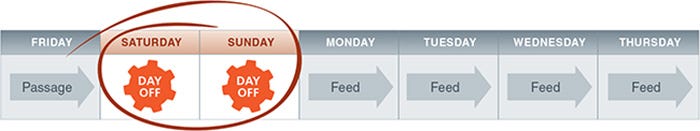
Figure 1. Overview of Weekend-Free Protocol
By following a routine 7-day passaging schedule on Fridays, weekend work is eliminated as medium replacement is not required on either Saturdays or Sundays.
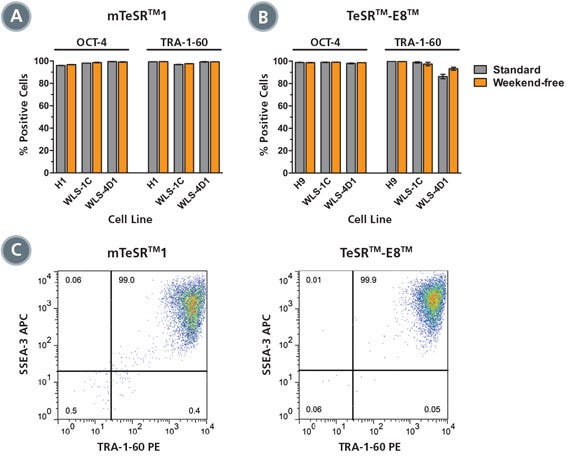
Figure 2. High Expression of Pluripotent Stem Cell Markers in Cells Cultured with the Weekend-Free Protocol
(A, B) Expression of pluripotent stem cell markers OCT-4 and TRA-1-60 was high after 10-11 passages for cells maintained in mTeSR™1 (A) or TeSR™-E8™ (B) using standard (gray) or weekend-free (orange) protocols (average of duplicate wells shown). (C) Representative FACS examples of weekend-free-cultured cells show co-staining of TRA-1-60 and SSEA-3 on nearly all cells (left: H1, mTeSR™1; right: H9, TeSR™-E8™).
Another important indicator of culture quality is the ability to maintain high rates of expansion during long-term culture without the acquisition of cytogenetic abnormalities. Cultures maintained without weekend feeding had similar expansion rates (Figure 4) and colony sizes to those following the standard 7-day passaging protocol with a weekend feed. This indicates that the reduced feeding schedule of the weekend-free 7-day passaging protocol does not impair the growth of the cultures. The cells also retained a normal karyotype (not shown) after long-term (> 10 passages) weekend-free maintenance in either mTeSR™1 or TeSR™-E8™.
Functional proof of differentiation to all three germ layers is ultimately required to verify retention of pluripotency after longterm passaging. We next assessed functional pluripotency of cells maintained with the standard or weekend-free 7-day passaging protocols using in vitro directed differentiation to cell types representing all three germ layers. Comparable efficiency of differentiation was achieved with cells from weekend-free cultures or standard cultures (Figure 5) for ectoderm using STEMdiff™ Neural Induction Medium (Catalog #05835), mesoderm using a cardiomyocyte differentiation protocol, and endoderm using the STEMdiff™ Definitive Endoderm Kit (Catalog #05110).
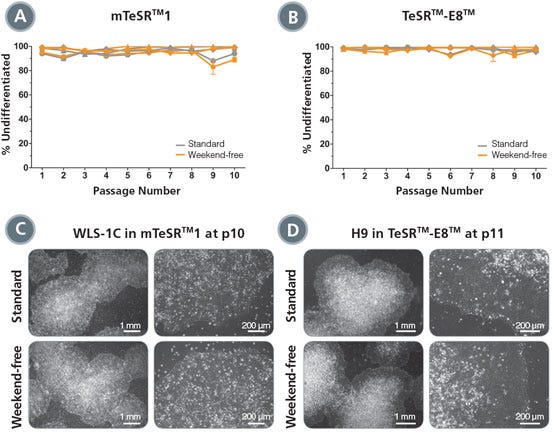
Figure 3. Normal Undifferentiated Colony Morphology is Maintained in Weekend-Free Cultures
(A, B) The percentage of colonies with undifferentiated morphology at each passage was determined by visual assessment of cultures of H1 or H9 (circles), WLS-1C (triangles) or WLS-4D1 (diamonds) cells maintained in mTeSR™1 (A) or TeSR™-E8™ (B) using standard (gray) or weekend-free (orange) protocols. All conditions showed consistently high percentages of undifferentiated colonies over 10 passages. (C, D) Representative images show comparable undifferentiated colony morphology in standard and weekend-free conditions.
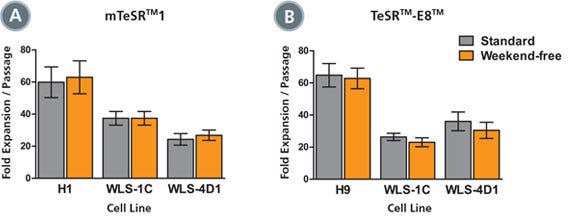
Figure 4. Expansion Rates of Weekend-Free Cultures are Comparable to Standard Cultures
Human ES or iPS cells were maintained in (A) mTeSR™1 or (B) TeSR™-E8™ using standard (gray) or weekend-free (orange) protocols. Fold expansion at each passage was determined by counting the number of aggregates harvested after 7 days of culture, divided by the number of aggregates seeded. Each bar represents the average of duplicate wells, averaged over 10 passages.

Figure 5. Ability to Differentiate to Three Germ Layers is Retained in Cells Cultured with the Weekend-Free Protocol
(A-C) Differentiation efficiency of WLS-4D1 cells previously maintained in mTeSR™1 for > 10 passages with standard or weekend-free protocols were comparable. (A) PAX6+ neural progenitor cells (red) with virtually no OCT-4+ undifferentiated cells (green) remaining (200X). (B) cTnT+ (cardiac troponin T) cardiomyocytes. (C) SOX17+CXCR4+ definitive endoderm cells.
Vitronectin XF™ Defined Matrix Can Be Used with the Weekend-Free Protocol
Vitronectin XF™ is a defined recombinant matrix protein that supports the growth of undifferentiated human ES and iPS cells. To test whether weekend-free culture could also be achieved using a completely defined system, H1 human ES cells were maintained using both the standard and weekend-free protocols, in mTeSR™1 and TeSR™-E8™, for five passages on Matrigel®, followed by five passages on Vitronectin XF™. We found that pluripotent cell marker expression, morphology, and expansion rates were equivalent during the first five passages on Matrigel® to those in the subsequent five passages on Vitronectin XF™ (Figure 6).
Cells cultured on either matrix using the weekend-free protocol had high expression of OCT-4 (Figure 6A), TRA-1-60, SSEA- 3, and SSEA-4 (not shown). Excellent morphology typical of highly undifferentiated human ES cells was also observed in all conditions (Figure 6B). The morphology of colonies on Vitronectin XF™ was characteristically more compact and rounded using either protocol (Figure 6C). A similar expansion rate was observed in all conditions (Figure 6D). Normal karyotype was confirmed after 10 passages without weekend feeding (Figure 6E).
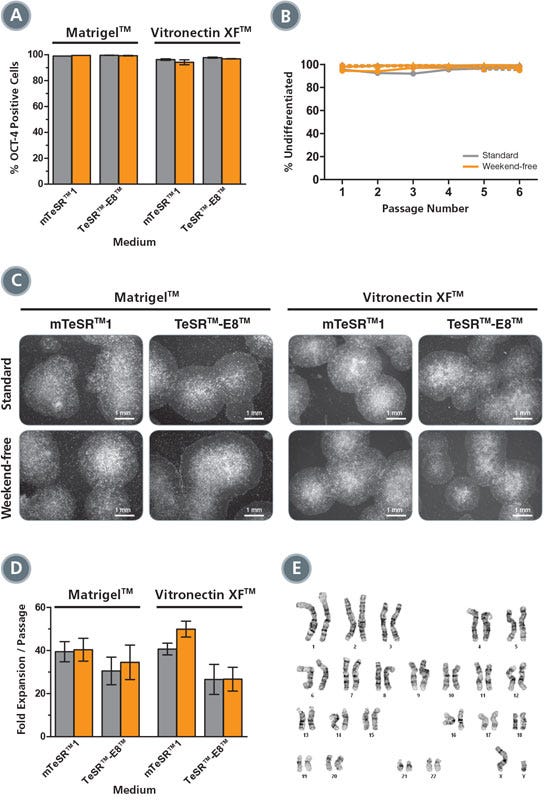
Figure 6. Vitronectin XF™ Supports Weekend-Free Maintenance Equivalently to Matrigel®
(A) After five passages on Matrigel® or Vitronectin XF™, virtually all cells expressed OCT-4 (average of duplicate wells). (B) Visual assessment at each passage showed equally high percentages of undifferentiated colonies on Matrigel® (solid lines) or Vitronectin XF™ (dashed lines), maintained in either mTeSR™1 (circles) or TeSR™-E8™ (triangles) using standard (gray) or weekend-free (orange) protocols. (C) Representative images of H1 colonies in all conditions show typical undifferentiated morphology. (D) Expansion rates were equivalent using standard (gray) or weekend-free (orange) protocols in all conditions (average of duplicate wells, averaged over 5 passages). (E) Normal karyotype of H1 cells cultured in mTeSR™1, after 10 passages without weekend feeding (5 passages Matrigel® plus 5 passages Vitronectin XF™).
Protocol Tips & Tricks
For details on general maintenance of human ES and iPS cells, without a strict 7-day passaging schedule, refer to the Technical Manuals ‘Maintenance of Human Pluripotent Stem Cells in mTeSR™1 and TeSR™2’ (Document #29106) or ‘Maintenance of Human Pluripotent Stem Cells in TeSR™-E8™’ (Document #29267). Successful adoption of a weekend-free protocol first requires mastering passaging at regular 7-day intervals. To convert a variable-day passaging schedule to a 7-day passaging schedule, there are two critical factors that must be optimized: 1) size of aggregates and 2) density of aggregates seeded.
Aggregate Size
- Colonies should be large and compact with dense centers at the time of passage (Day 7, Figure 7D).
- Aggregate and colony size can be measured using an eyepiece micrometer (also called a reticule or graticule), calibrated against a standard micrometer.
- Break up aggregates by gently pipetting up and down, to a relatively homogeneous size of about 80 - 150 μm in diameter (Figure 7A; yellow arrows).
- If aggregates are too large (> 200 μm), colonies will be ready for passage prior to day 7, and if left until the 7th day, will become too dense and overtly differentiated.
- If aggregates are too small (< 60 μm), they may attach poorly to the matrix and colonies will not be ready for passage on day 7 (Figure 7A; red arrow heads).
- When switching to a 7-day passaging protocol, monitor colony size to confirm the seeded aggregate size was
- Day 1: colonies should be quite small, about 100 - 300 μm in diameter, or 20 - 200 cells (Figure 7B).
- Day 3: colonies should be approximately 300 - 700 μm in diameter with few over 1 mm (Figure 7C).
- Day 7: colonies should be tightly packed with dense centers and reach 2 - 4 mm in diameter (Figure 7D).
Seeding Density
- Cultures should be 70 - 90% confluent at the time of passage.
- To attain consistent seeding density, cell aggregates should be counted and plated in defined numbers.
- To determine aggregate concentration, place an aliquot of the aggregate suspension in a flat-bottom 96-well plate and count all aggregates ≥ 60 μm in diameter.
- Seed optimized number of aggregates into new wells. Recommended range ~30 - 50 aggregates/cm2 (e.g. 300 - 500 aggregates/well of a 6-well plate).
- Adjust this number as necessary to achieve the correct confluency at day 7. Optimal seeding density can vary depending on factors such as cell line, but should be consistent at each passage.
- If too many aggregates are seeded, cultures will become overly confluent by day 7 and show signs of differentiation.
- If too few aggregates are seeded, cell yields will be lower.
- This seeding density typically results in a split ratio of between 1 in 10 and 1 in 40.
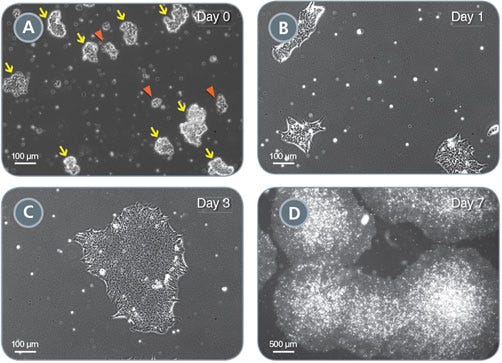
Figure 7. Colony Growth During 7-Day Culture
(A) Aggregate suspension containing optimally-sized aggregates (80 - 150 μm in diameter; yellow arrows). Single cells and aggregates of < 60 μm in diameter (red arrowheads) should be minimized. (B) Representative day 1 colonies after a medium change. (C) Representative day 3 colony. (D) Representative day 7 colonies ready for passaging.
Summary
By manipulating aggregate size, culture density and passaging day, it is possible to maintain human ES or iPS cell cultures without working on the weekend. This requires the plating of small cell aggregates at low densities to establish a regular 7-day passaging schedule on Fridays. Once this routine is established, cultures can tolerate two full days (the weekend) without feeding, and cells from these cultures will maintain undifferentiated morphology, pluripotent cell marker expression and high expansion rates.
Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration

