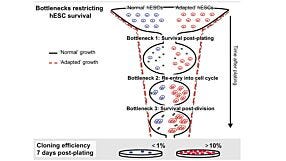
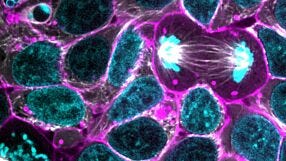
Re-Creating Disease with Kidney Organoids and CRISPR
In this webinar, Dr. Benjamin Freedman from the University of Washington provides an overview of re-creating disease using kidney organoids and CRISPR-Cas9 genome editing. As a postdoctoral fellow in the Renal Division at Brigham and Women’s Hospital at Harvard Medical School, Dr. Freedman was responsible for several innovative techniques that he describes in this webinar, including differentiation of human pluripotent stem cells (hPSCs) into kidney organoids containing nephron-like segments and generating knockout mutations in organoids with CRISPR.
In this webinar, Dr. Benjamin Freedman from the University of Washington provides an overview of re-creating disease using kidney organoids and CRISPR-Cas9 genome editing. As a postdoctoral fellow in the Renal Division at Brigham and Women’s Hospital at Harvard Medical School, Dr. Freedman was responsible for several innovative techniques that he describes in this webinar, including differentiation of human pluripotent stem cells (hPSCs) into kidney organoids containing nephron-like segments and generating knockout mutations in organoids with CRISPR.
Webinar Q&A
Below is a list of all questions submitted by participants during the live viewing of the webinar. Answers are provided by Dr. Benjamin Freedman. 1. Are your whole organoid IF pictures from 3D whole mount staining or from sections of the organoids? Whole mount. This is relatively easy to do in our differentiation system. 2. Which marker do you use for the endothelial network? CD31 is very reliable and specific. There are other equally good ones. 3. It is thought that endothelial cells migrate to embryonic kidney form surrounding tissue. How would you interpret where the endothelial cells in organoids come/differentiate from? Do they differentiate in response to CHIR or NPS niche signals? CHIR is used in a variety of endothelial differentiation protocols, so it could be doing the same thing here. There may also be niche signals, since it is clear that the endothelial cells are attracted to the organoids (or vice versa). 4. Does CRISPR modified iPSCs change gene expression due to freeze thaw cycle? I am sure that freeze thawing has some effect on gene expression! We always isolate unmodified controls alongside our CRISPR mutants, from the same transfections, and treat them the same way. 5. In the PC1/2 IF pictures looking at the co-localisation there seems to be a lot of background on each channel with just the two bright spots - is this unavoidable? And how can you be sure the bright points are not just artifacts? The PC2 antibody we use does give some background, and we see it also in the PKD2-/- mutants. We know it's artifact because it's still there in the stainings but the protein is completely gone by Western blot. 6. Did you stain the orgaoids for presence of primary cilia? The organoids and cysts possess abundant primary cilia, as shown in our 2017 Nature Materials paper. But, we didn't do an exhaustive characterization. 7. Which method (NHEJ or HDR) was used to generate PKD1 -/- and PKD2 -/- ? Would the choice of method have any consequences for cystogenesis? NHEJ was used, since we just wanted to make knockouts. HDR could also be used to introduce specific nonsense mutations, which would presumably lead to the same phenotype, but we haven't done that yet. 8. Is there a way to introduce vasculature during kidney organoid differentiation from these mutant cells? There are endothelial cells in the cultures, but we have not yet investigated whether these have PKD phenotypes. Would be interesting to do so, since PKD patients are at risk for aneurysms. 9. What is the rationale behind placing detached cysts into in vitro cystogenic conditions? Could the impressive size difference of the PKD mutant cysts vs control be explained by increased proliferation only? I am not sure what the reference is to 'cystogenic conditions.' We place organoids containing tubules into 3D cultures. The PKD organoids form cysts, the controls do not. Proliferation plays a major role but most of the difference in size is probably due to fluid accumulation inside the cyst. 10. PKD2 in your model is implicated in adhesion to substrate. Do you propose that mechanistically it acts at the primary cilia site or rest of plasma membrane? Hard to say at this point. If the polycystins are directly mediating adhesion, it's not quite clear how cilia could be doing that. 11. Are you looking into any other functional tests for the organoids other than Cisplatin and Gentamicin assay, and dextran uptake? There are many functional tests that would be interesting to explore in these organoids. Nephrotoxicity and dextran uptake work well so we have focused on those. There are others we have tried and not published because they haven't worked. 12. Did you have ever notice if these cystic organoid/cysts also having air filled inside that helps to float them in culture when comes out from their organoid? If there was air we'd see bubbles, we don't see that. 13. Is there any report to introduce parallel nerve cell or endothelial cell differentiation within these organoids? See our 2015 Nature Communications paper. Both nerves and endothelial cells are already present within these cultures. 14. With the advent of 3D printing, do you envision using other types of matrices, other than Matrigel, etc.? It would certainly be interesting to examine a range of natural and synthetic matrices, to see how they affect cyst formation. 15. Did you ever try to combine the kidney organoids with microfluidics to achieve filtration in the tubules? We are working on microfluidics, but I'm not sure it will be easy to achieve filtration without an integrated vasculature. The tubules do not have obvious inlets and outlets that can be perfused. 16. What is the meaning of the suspension culture? Why do you think suspension culture is quite important to get the readout? How do you think to use this method? Suspension culture means that the organoids have been released from their adherent cues. This allows the organoids to take on their more natural shape. We thought of this because we noticed that cyst formation in the adherent cultures involved detachment of the tubules from the plastic. Also, we were familiar with suspension culture because we use it to make embryoid bodies to test the general quality of our iPS cell lines. 17. Following genome editing, did you test for any off-target effects, and if so what strategy did you use? It's very hard to identify off-target effects directly, but those who have done this in the past have shown that they are uncommon in ES cells. Since we get the same phenotype with multiple guide RNAs, and multiple clones, we don't think the phenotypes we observe are due to off-target effects. Targeting other genes produces different phenotypes. 18. How important do you think vascularization will be for your organoid model in the future, and do you think this will be achievable? It would be great to get a bona fide vasculature, but it is hard to say how achievable it will be in organoids. It is clear that the endothelial cells in our organoids don't form bona fide glomeruli, even though they are adjacent to podocytes, so there is clearly something missing. Probably we need to start in vivo and see if we can get that to work before moving in vitro.Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration