Gating Strategy for the Identification of ILC2s
- Document # 27030
- Version 2.1.0
- Aug 2020
Background
Studies starting in 2010, led several research groups to identify a small population of immune cells that have characteristics of lymphoid cells but lack lineage markers (Lin-) and re-arranged antigen specific cell surface receptors.1,2 As these cells were identified they were referred to by different names such as natural helper cells,3,4 nuocytes,5 innate lymphoid cells6 and innate helper 2 cells.7 In 2013 a uniform nomenclature was proposed to classify these cells as group 2 innate lymphoid cells (ILC2s).8 ILC2s are part of the larger ILC family, that according to this new nomenclature, is divided into three major groups: group 1 ILCs (ILC1s), group 2 ILCs (ILC2s) and group 3 ILCs (ILC3s).8 The groupings are based on transcription factor expression, surface receptors and cytokine-secreting profiles that mirror the profiles of T helper subsets, Th1, Th2 and Th17 respectively. Accordingly, ILC2s produce Th2 associated cytokines including interleukin (IL)-4, IL-5, IL-9 and IL-13 and are involved in type 2 immune response (Figure 1). By surface staining, human ILC2s are negative for lineage markers and stain positive for CD45, CD294, CD127 and CD161. Mouse ILC2s are negative for lineage markers and positive for CD45, ICOS and ST2 (at varying levels). Since their initial identification, we now know that ILC2s are important effector cells implicated in innate immunity and are required for anti-helminth immunity, allergic inflammation and tissue repair.1 More information on the development, biology and function of the distinct ILC subsets, including ILC2s, can be found in several publications.1,2,9,10
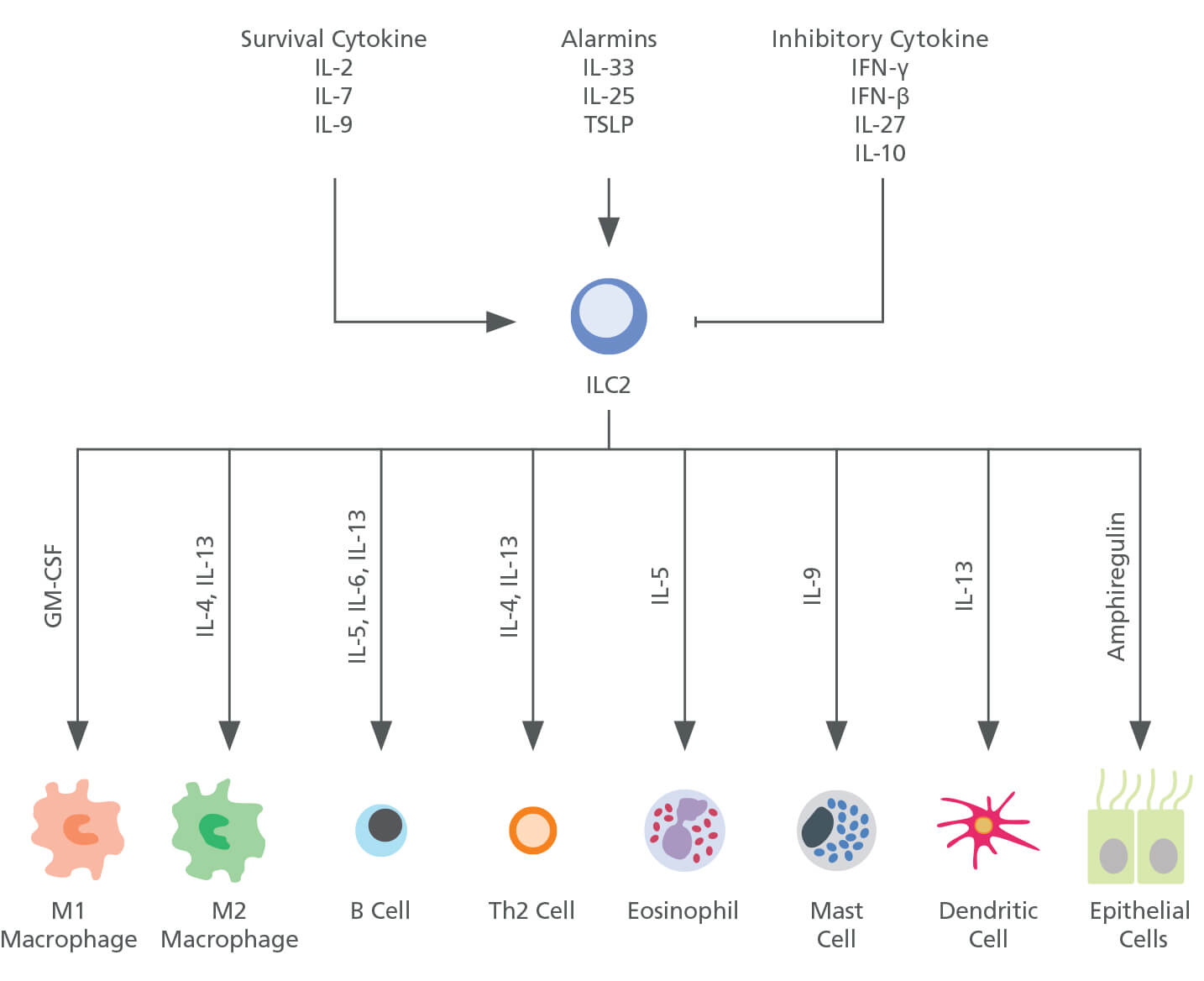
Figure 1. Group 2 Innate Lymphoid Cells (ILC2s)
ILC2s respond to a diverse range of stimuli including survival cytokines, such as IL-2, IL-7 and IL-9, and alarmins such as IL-33, IL-25 and TSLP. ILC2s then produce type-2 cytokines, including IL-4, IL-5, IL-9 and IL-13 and thereby promote activation of macrophages, B cells, CD4+ T cells, basophils, eosinophils, mast cells, dendritic cells and epithelial cells. GM-CSF, granulocyte macrophage colony-stimulating factor; IFNγ, interferon-γ; IL, interleukin; TSLP, thymic stromal lymphopoietin.
The Challenge
ILC2s are widely distributed throughout the body. They are enriched at barrier surfaces such as the skin, intestine and lung, but are also present in adipose tissue and peripheral blood.1 Even though ILC2s are found in different tissues they comprise less than 1% of the total leukocyte population in human and mice.11,12 The low frequency and lack of specific surface markers for ILC2s and ILCs in general, creates challenges when it comes to identification, isolation and further characterization of these cells. Currently, ILC2s are isolated using multicolor flow cytometric cell sorting but this method is time-consuming, expensive and can result in low cell recovery. To address this challenge, STEMCELL Technologies has developed products to enrich for ILC2 populations from human and mouse tissue using the RosetteSep™ and EasySep™ technology (see page 4). Pre-enrichment of ILC2s will reduce sorting time and improve purity and recovery. The enriched ILC2s can then be further prepared for analysis or sorting using flow cytometry.
Gating Strategy for the Identification of Human and Mouse ILC2s
Cell Pre-Enrichment and Antibody Labeling
Traditionally, to sort or analyze ILC2s by flow cytometry, one would begin by collecting the tissue of interest and prepare a single cell suspension using a suitable protocol. The entire sample is then labeled with appropriate fluorochrome-conjugated antibodies and a viability dye. Recommended antibodies for labeling human and mouse cells are shown in Table 1 and Table 2, respectively. Note that both human and mouse ILC2s are defined as lineage-negative (Lin-) cells. In order to gate the Lin- cell population more easily, ensure that all antibodies recognizing lineage markers have the same fluorochrome (e.g. FITC). The user will then proceed to a flow cytometer and continue with the analysis or sorting of rare ILC2s.
Since ILC2s are present at a low frequency, we recommend performing a pre-enrichment step using EasySep™ or RosetteSep™ ILC2 enrichment kits (see page 4) before labeling cells with fluorochrome-conjugated antibodies. By removing the bulk of unwanted cells and pre-enriching for ILC2s, the user is able to process their sample more quickly while improving the recovery of ILC2s on a flow cytometer.
Identification of Human ILC2s from Blood
A representative gating strategy to identify human ILC2s from unenriched and enriched human whole blood samples is shown in Figure 2. The same gating strategy can be applied when analyzing cells from leukapheresis samples (not shown). Following the gating strategy shown on Figure 2, start on the SSC-A versus FSC-A plots and create a gate to include all leukocytes based on cell granularity and cell size. Alternatively, create a smaller gate around only the lymphocytes (not shown). When sorting cells, remove doublets and gate on single cells (singlets) using either the FSC-W versus FSC-H or SSC-W versus SSC-H plots (not shown). Then, select for viable cells and then for CD45+ cells. Next, create a gate on the Lin- CRTH2+ (CD294) population. Within this subset the ILC2 population is positive for CD127 and CD161..
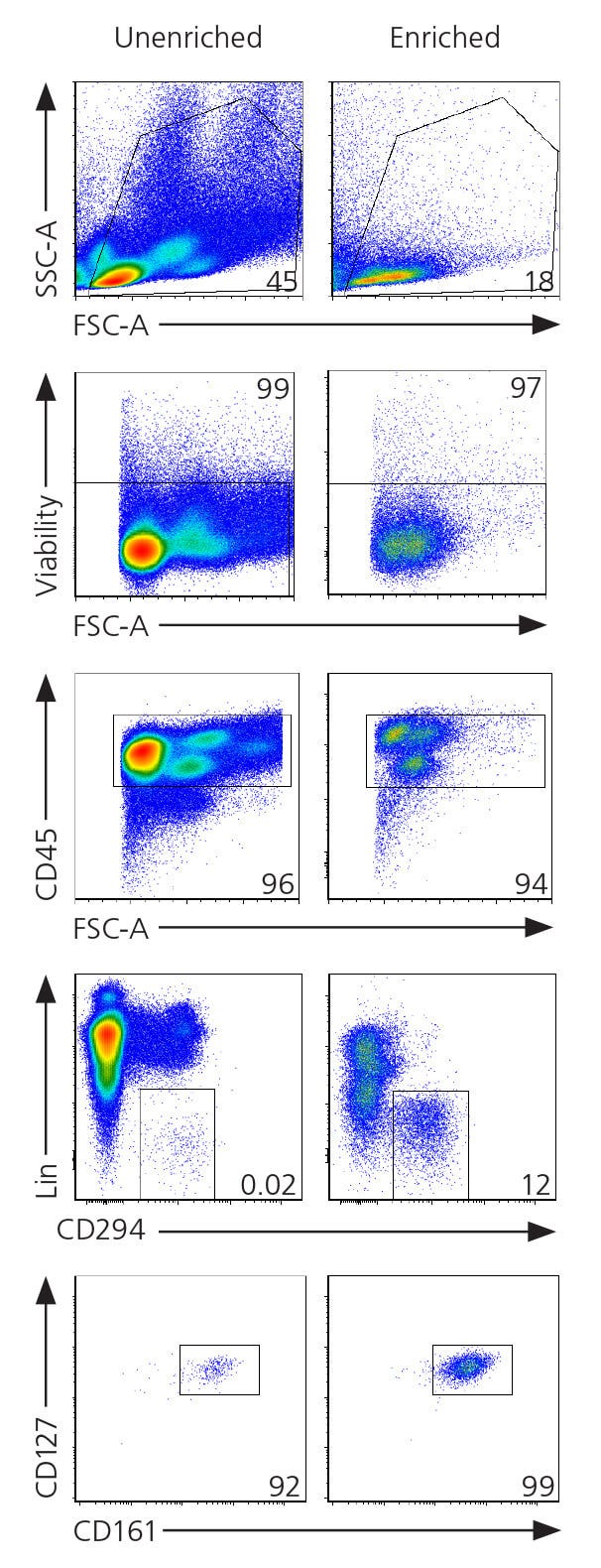
Figure 2. Gating Strategy for ILC2s From Human Blood Samples
Enriched cells were obtained using RosetteSep™ Human ILC2 Enrichment Kit (Catalog #15382) as shown in page 4. For the unenriched sample, cells were isolated from human whole peripheral blood using density gradient centrifugation. The frequency of ILC2s (Lin-CD45+ CD294+CD127+ CD161+) in the enriched fraction typically ranges from 0.44 - 53% and is donor dependent. In this example, the percentage of ILC2s in the unenriched and enriched fractions corresponds to 0.02 and 12%, respectively. FACS plots were generated using FCS Express 5 Software (De Novo Software).
Table 1. Antibodies for Flow Cytometric Analysis and Cell Sorting of Human ILC2s
Identification of Mouse ILC2s from Lung
A representative gating strategy to identify mouse ILC2s from unenriched and enriched single-cell suspension from lungs is shown in Figure 3. The same gating strategy may be applied for cells from other tissues. However, the starting frequency and cell types will vary between different tissues. Therefore, ILC2s may have a somewhat different profile and the plots may look slightly different from what is shown in Figure 3. Start on the SSC-A versus FSC-A plots and create a gate to include all leukocytes. Alternatively, create a smaller gate around the lymphocytes only (not shown). When sorting cells, remove doublets and gate on single cells (singlets) using either the FSC-W versus FSC-H or SSC-W versus SSC-H plots. Then, select for viable cells and CD45+ cells. Next, create a gate on the Lin- ICOS+ population. Within this subset the ILC2 population is positive for CD90 and ST2. Mouse ILC2s from the lungs are described as Lin-, CD45+, ICOS+, ST2+, CD90+. ILC2s are also positive for transcription factor, GATA-3. Other markers commonly used to identify mouse ILC2s include KLRG1, CD127 and c-Kit.3,4
Intranasal administration of IL-33 in mice has been shown to increase the frequency of ILC2s found in the lung.12 To get familiar with ILC2 enrichment and sorting, IL-33 treated lungs can be used as positive control (Figure 4).
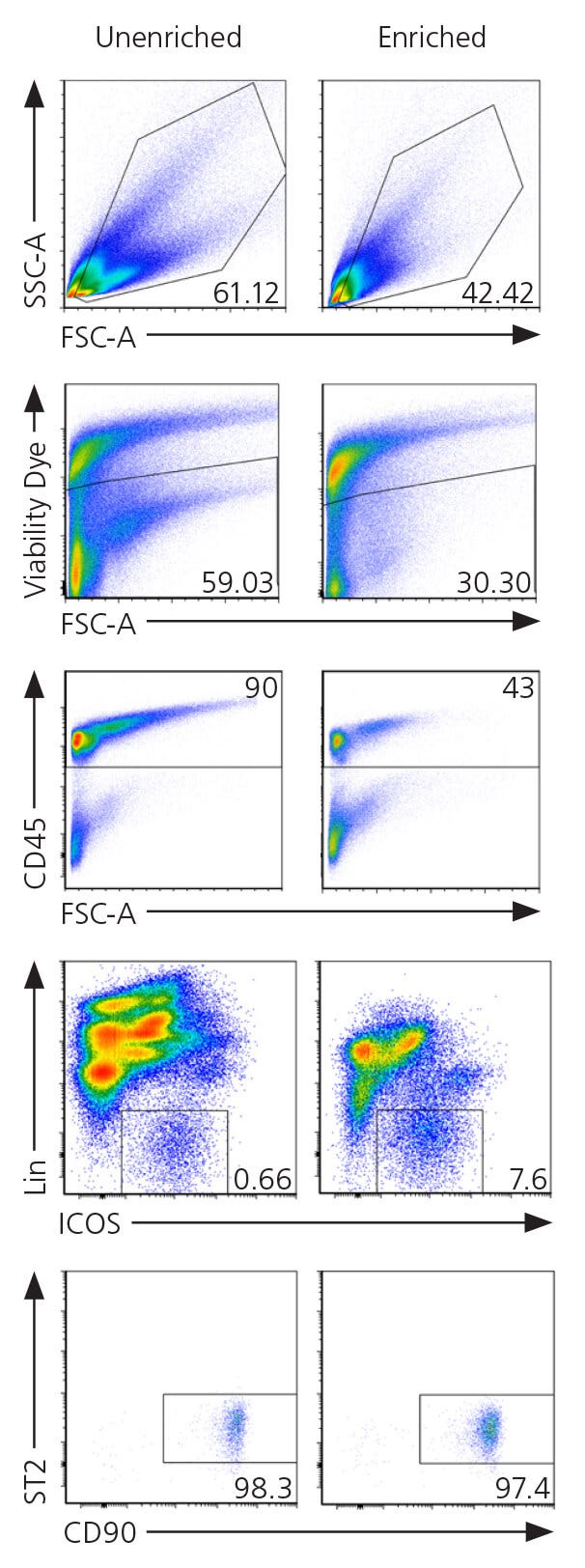
Figure 3. Gating Strategy for ILC2s From Mouse Lungs
Single cell suspension from C57Bl/6J mouse lung tissue were enriched using EasySep™ Mouse ILC2 Enrichment Kit (Catalog #19842) as shown in page 4, or left unenriched. The ILC2 content (Lin-CD45+ ICOS+ST2+) in the enriched fraction typically ranges from 2.2 - 7.1%. In this example, the percentage of ILC2s in the unenriched and enriched fractions correspond to 0.8% and 16.65% of CD45+ cells respectively. FACS plots were generated using FCS Express 5 Software (De Novo Software).
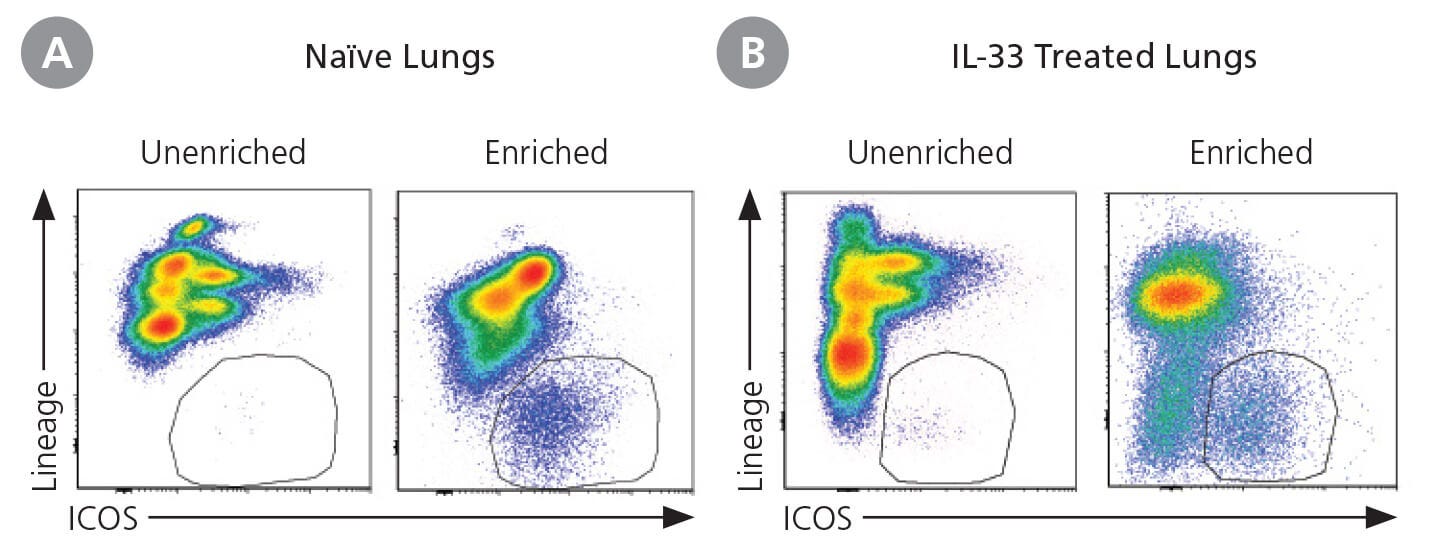
Figure 4. ILC2s From Lungs From Naive and IL-33 Treated Mice
ILC2s were isolated from the lungs of naïve or IL-33 treated C57Bl/6J mice as previously described.13 Multiple lungs were pooled (unenriched sample) and a portion of the pooled sample was then enriched using EasySep™ Mouse ILC2 Enrichment Kit (Catalog #19842).
Table 2. Antibodies for Flow Cytometric Analysis and Cell Sorting of Mouse ILC2s
Advantages of ILC2 Enrichment Before Sorting
- Enrich cells without the need for columns by using EasySep™ or RosetteSep™.
- Reduce the time spent sorting cells.
- Improve the purity and yield of sorted ILC2s.
EasySep™ Protocol
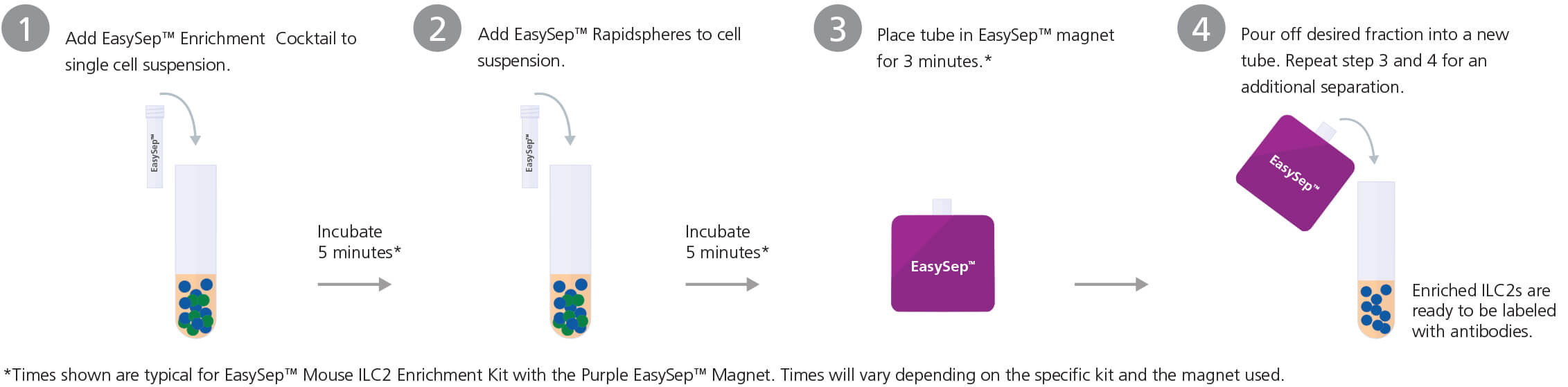
RosetteSep™ Protocol
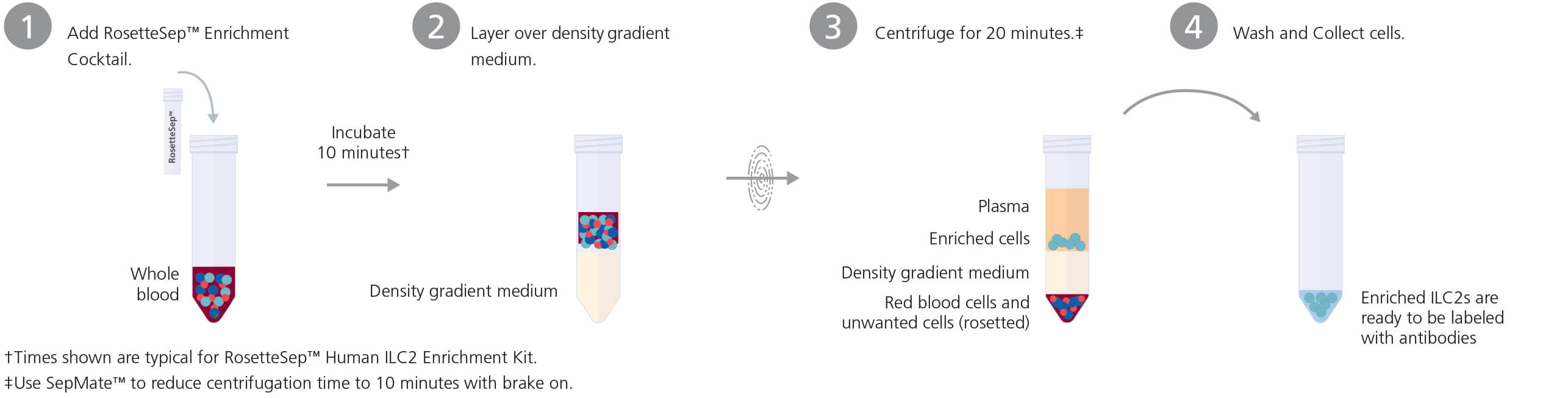

Product Listing
References
- Artis D & Spits H. (2015) The biology of innate lymphoid cells. Nature 517(7534): 293–301.
- Spits H & Cupedo T. (2012) Innate lymphoid cells: emerging insights in development, lineage relationships, and function. Annu Rev Immunol 30(1): 647–75.
- Moro K et al. (2010) Innate production of T(H)2 cytokines by adipose tissue-associated c-Kit(+)Sca-1(+) lymphoid cells. Nature 463(7280): 540–4.
- Halim TYF et al. (2012) Lung natural helper cells are a critical source of Th2 cell-type cytokines in protease allergen-induced airway inflammation. Immunity 36(3): 451–63.
- Neill DR et al. (2010) Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464(7293): 1367–70.
- Buonocore S et al. (2010) Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature 464(7293): 1371–5.
- Price AE et al. (2010) Systemically dispersed innate IL-13-expressing cells in type 2 immunity. Proc Natl Acad Sci U S A 107(25): 11489–94.
- Spits H et al. (2013) Innate lymphoid cells--a proposal for uniform nomenclature. Nat Rev Immunol 13(2): 145–9.
- Walker JA et al. (2013) Innate lymphoid cells--how did we miss them? Nat Rev Immunol 13(2): 75–87.
- Sonnenberg GF & Artis D. (2012) Innate lymphoid cell interactions with microbiota: implications for intestinal health and disease. Immunity 37(4): 601–10.
- Mjösberg JM et al. (2011) Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nat Immunol 12(11): 1055–62.
- Halim TYF & Takei F. (2014) Isolation and characterization of mouse innate lymphoid cells. Curr Protoc Immunol 106(5): 3.25.1-13.
- Martinez-Gonzalez I et al. (2016) Allergen-Experienced Group 2 Innate Lymphoid Cells Acquire Memory-like Properties and Enhance Allergic Lung Inflammation. Immunity 45(1): 198–208.
Request Pricing
Thank you for your interest in this product. Please provide us with your contact information and your local representative will contact you with a customized quote. Where appropriate, they can also assist you with a(n):
Estimated delivery time for your area
Product sample or exclusive offer
In-lab demonstration

