Studying COVID-19 with Intestinal Organoids
Why Are Intestinal Organoids a Relevant Model for Viral Infection?
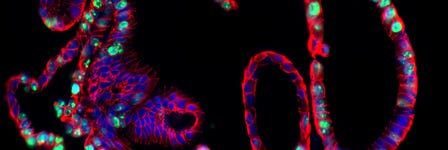
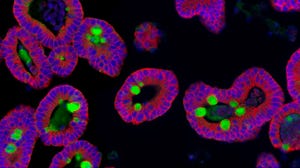
COVID-19 patients frequently experience gastrointestinal symptoms and, consequently, there has been some investigation into the nature of viral infection in the intestine.1, 2 Intestinal organoids express factors critical for SARS-CoV-2 entry into cells2, 3 and are valuable in vitro models to understand coronavirus infection in the gastrointestinal tract.
Although SARS-CoV-2 infection most obviously causes respiratory symptoms, it is also often characterized by multi-organ involvement. The viral receptor protein angiotensin-converting enzyme 2 (ACE2)4 is expressed in several epithelial tissues, including the intestine, alongside TMPRSS2, a protease that activates the viral spike protein for binding.3
To investigate the molecular mechanisms of SARS-CoV-2 infection in the intestine, researchers are using intestinal organoids, which recapitulate in vitro the cellular identity and molecular characteristics of the intestinal epithelium.1, 2 Intestinal organoids provide a useful model for SARS-CoV-2 invasion and transmission studies, as they can be rapidly expanded and maintained and are amenable to a wide range of in vitro manipulations. Organoid-derived monolayers are also convenient models for viral invasion, as they enable direct access to the apical epithelial surface.
Key Publications
Lamer MM et al. (2020) SARS-CoV-2 productively infects human gut enterocytes. Science: eabc1669. Epub ahead of print, DOI: 10.1126/science.abc1669.
In this paper, researchers from the Clevers and Haagmans labs in the Netherlands use human small intestinal organoids as a model for infection and replication of SARS-CoV-2 virus in vitro. Both SARS-CoV and SARS-CoV-2 were shown to infect intestinal organoids and produce infectious progeny.
Zang R et al. (2020) TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci Immunol 5(47):eabc3582.
Researchers used healthy human small and large intestinal organoids to assess the ability of SARS-CoV-2 to infect and replicate inside intestinal epithelial cells. They investigated the presence of ACE2 and TMPRSSs in both three-dimensional organoids and organoid-derived monolayers and used both CRISPR-based gene-edited organoids and therapeutic serine protease inhibition to test the contributions of TMPRSS2 and TMPRSS4 to infection.
Intestinal Organoids in COVID-19 Research
The emergence of intestinal organoid technologies a decade ago5, 6 provided researchers with a mode for in vitro investigation of intestinal epithelial tissue. Intestinal organoids recreate in vitro the cellular complement of the intestinal epithelium and retain the genotype and phenotype of the parental cells; in many respects, they provide a faithful recapitulation of the in vivo tissue. This methodology opened up new experimental approaches that were previously not possible with in vivo experiments, immortalized cell lines, or short-term cultures of primary intestinal cells.
In addition to expressing the SARS-CoV-2 viral receptor protein, ACE24, at the apical epithelial surface, intestinal organoids have been shown to be readily infected by the SARS-CoV-2 virus, which results in infectious progeny.1, 2 They thus provide a model system for studying the pathogenesis of SARS-CoV-2, both in a basic research context as well as during the development of therapeutic agents to combat COVID-19. Explore the publication list in the next section below to see how intestinal organoids have been used to study other infectious agents that infect the intestine.
While intestinal organoids are typically polarized such that their apical surface is on the interior/luminal surface of the organoid, growing them as a 2D organoid-derived monolayer enables easy access to both the apical and basolateral surfaces. This modification of the organoid culture system facilitates studying the interaction of viral particles, including SARS-CoV-22, and bacterial cells with the intestinal epithelium. These organoid-derived monolayers can be air-lifted at the air-liquid interface (ALI) to achieve further differentiation of the epithelium. ACE2 expression is observed in organoid-derived monolayers, with strong localization to the apical surface (Figure 1).
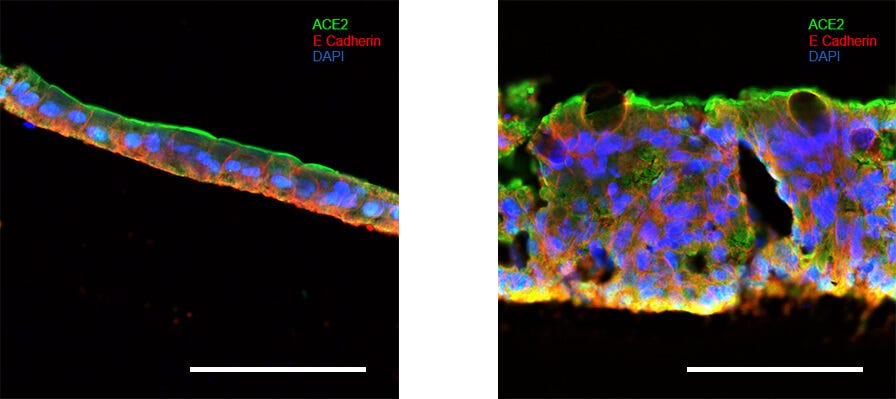
Figure 1. Intestinal Organoid-Derived Monolayers Express ACE2 at Their Apical Surface
Intestinal organoids were grown in IntestiCult™ Organoid Growth Medium (Human) and used to seed organoid-derived monolayers. Immunofluorescence staining of (Left) organoid-derived submerged monolayers and (Right) organoid-derived monolayers cultured at the ALI show strong staining for ACE2 at their apical surface. Scale bars = 100 μm.
Infectious Disease Studies Using IntestiCult™
Coronavirus (CoV)
- Li L et al. (2019) Porcine intestinal enteroids: a new model for studying enteric coronavirus porcine epidemic diarrhea virus infection and the host innate response. J Virol 93(5): e01682–18.
- Li L et al. (2019) IFN-lambda 3 mediates antiviral protection against porcine epidemic diarrhea virus by inducing a distinct antiviral transcript profile in porcine intestinal epithelia. Front Immunol 10: 2394.
Norovirus (NoVs)
- Ford-Siltz LA et al. (2020) Genotype-specific neutralization of norovirus is mediated by antibodies against the protruding domain of the major capsid protein. J Infect Dis jiaa116. Epub ahead of print, DOI: 10.1093/infdis/jiaa116.
- Altmar RL et al. (2020) Comparison of microneutralization and histo-blood group antigen-blocking assays for functional norovirus antibody detection. J Infect Dis 221(5): 739–743.
- Haga K et al. (2020) Genetic manipulation of human intestinal enteroids demonstrates the necessity of a functional fucosyltransferase 2 gene for secretor-dependent human norovirus infection. mBio. 11(2): e00251–20.
Escherichia coli
- Yahiro K et al. (2018) Mechanism of inhibition of Shiga-toxigenic Escherichia coli SubAB cytotoxicity by steroids and diacylglycerol analogues. Cell Death Discov 4: 22.
How Are Intestinal Organoids Grown?
Intestinal organoids can be cultured either by directed differentiation of human pluripotent stem cells (hPSCs) or from intestinal stem cells (also known as adult or tissue-resident stem cells) isolated from the crypt base of intestinal or colonic primary tissue samples. These lineage-restricted stem cells are responsible for continual regeneration of the intestinal epithelium in vivo.
To form organoids from intestinal stem cells, intestinal crypts are embedded in an extracellular matrix and cultured in a medium that incorporates signaling factors to mimic the environment of the in vivo intestinal stem cell niche. This environment, which includes both physical and chemical signaling, allows the intestinal stem cell population to expand and give rise to the range of progeny cells that form the complement of intestinal epithelial cells in vivo.
When starting with hPSCs, the cells are first directed to differentiate to the endoderm and then mid-/hindgut lineage, at which point spheroids spontaneously bud off of the culture monolayer and are collected and embedded in extracellular matrix, similar to the method described above. In comparison to intestinal stem cell-derived organoids, hPSC-derived intestinal organoids incorporate mesenchymal cells and are much more fetal in nature.7
Organoid-Derived Monolayer and ALI Cultures
While intestinal organoids can be readily expanded and maintained in long-term culture, the apical surface of the cells is oriented toward the organoid lumen, which limits experimental access to the side of the epithelium that is exposed to foreign material in vivo. Plating dissociated intestinal stem cell-derived organoids as monolayers generates cultures that provide easy access to the apical surface.
Compared with expanding organoids, these organoid-derived monolayers exhibit more extensive differentiation of the intestinal cells and this can be enhanced even further by air-lifting the cultures to grow them at the ALI. These differentiated cultures closely mimic the mature intestinal epithelium and enable convenient investigation of viral invasion and other interactions at both the apical and basal surfaces of the epithelium.
The defining feature of ALI culture is that the basal surface of the cells is in contact with liquid culture medium, whereas the apical surface is exposed to air. A common approach is to seed cells onto the permeable membrane of a cell culture insert, which is initially supplied with culture medium in both the apical and basal compartments (Figure 2A). Once confluence is reached, the cells are subjected to ‘air-lift’, where the medium is removed from the apical chamber and supplied only to the basal chamber (Figure 2B).
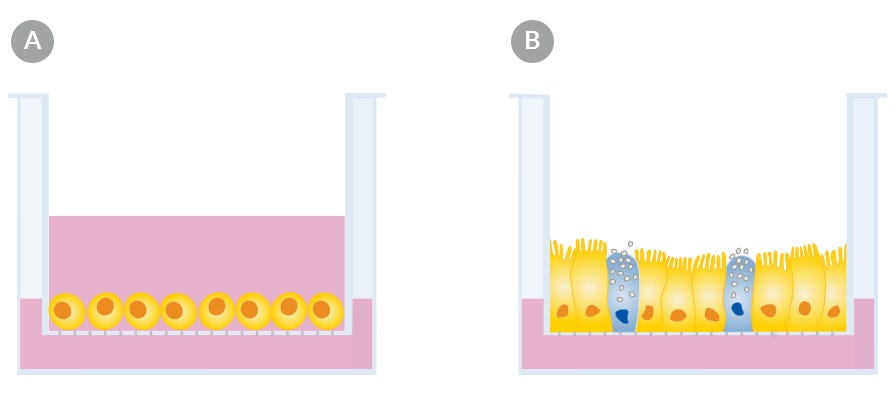
Figure 2. Air-Liquid Interface Culture
Culture at the air-liquid interface drives further differentiation of intestinal organoid-derived monolayers. (A) Monolayers are established and allowed to reach confluence. (B) Once cultures reach confluence, the medium is removed from the apical surface while medium is still supplied to the basolateral surface.
Products for COVID-19 Research
IntestiCult™ Organoid Growth Medium (Human)
Establish and maintain human intestinal organoids from intestinal stem cells with this cell culture medium.
View all products for intestinal epithelial cell research >
Learn more about airway ALI and kidney organoid models for COVID-19 research.
More Tools for Organoid Research
Related Resources
Nature Research Roundtable: Organoids
Learn key insights from the discussions and watch the presentations from our Nature Research Roundtable on organoids.
Organoid E-Book
Read the Wiley "Essential Knowledge Briefing" on the evolution and applications of organoid research.
Patient-Derived Organoids for Drug Screening and Development
Dr. Sylvia Boj describes how patient-derived intestinal organoids are being used to screen cystic fibrosis patients for treatment response.
References
- Lamers MM et al. (2020) SARS-CoV-2 productively infects human gut enterocytes. Science eabc1669. Epub ahead of print, DOI: 10.1126/science.abc1669.
- Zang R et al. (2020) TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci Immunol 5(47): eabc3582.
- Sungnak W et al. (2020) SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat Med 26(5): 681–687.
- Zhou P et al. (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798): 270–273.
- Sato T et al. (2009) Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459(7244): 262–5.
- Sato T et al. (2011) Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett's epithelium. Gastroenterology 141(5): 1762–72.
- Kretzmar K and Clevers H (2016) Organoids: Modeling development and the stem cell niche in a dish. Dev Cell 38(6): 590–600.